- Mathematics

The Effect of Computer Laboratory Facilities and Learning Interest on Students’ Learning Outcomes
- Kreano Jurnal Matematika Kreatif-Inovatif 12(1):97-106
- 12(1):97-106
- CC BY-NC-SA 4.0

- Universitas Islam Negeri Alauddin

- Institut bisnis dan teknologi pelita indonesia
Abstract and Figures

Discover the world's research
- 25+ million members
- 160+ million publication pages
- 2.3+ billion citations
- Ardhi Prabowo
- Amidi Amidi
- Detalia Noriza Munahefi
- Tuan Van Nguyen
- Hung Thanh Nguyen
- Chinh Danh Cao
- Hang Thi Thuy Vu
- Agustin Hanivia Cindy
- Sugiyono Sugiyono
- Husaini Usman

- Ana Muliyana
- Adi Wibowo Panjaitan
- Frengki Simatupang

- Nelly Budiyarti
- Roida Eva Flora Siagian
- Zakiyatul Lutfiyah

- Y B Kusumah
- Recruit researchers
- Join for free
- Login Email Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google Welcome back! Please log in. Email · Hint Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google No account? Sign up
- Skip to Content
- Bulletin Home

- This Is MIT >
- Research and Study >
Computer Science and Artificial Intelligence Laboratory
- Around Campus
- Academic Program
- Administration
- Arts at MIT
- Campus Media
- Fraternities, Sororities, and Independent Living Groups
- Medical Services
- Priscilla King Gray Public Service Center
- Religious Organizations
- Student Government
- Work/Life and Family Resources
- Advising and Support
- Digital Learning
- Disability and Access Services
- Information Systems and Technology
- Student Financial Services
- Writing and Communication Center
- Major Course of Study
- General Institute Requirements
- Independent Activites Period
- Undergraduate Research Opportunities Program
- First-Year Advising Seminars
- Interphase EDGE/x
- Edgerton Center
- Grading Options
- Study at Other Universities
- Internships Abroad
- Career Advising and Professional Development
- Teacher Licensure and Education
- ROTC Programs
- Financial Aid
- Medical Requirements
- Graduate Study at MIT
- General Degree Requirements
- Other Institutions
- Registration
- Term Regulations and Examination Policies
- Academic Performance and Grades
- Policies and Procedures
- Privacy of Student Records
- Abdul Latif Jameel Poverty Action Lab
- Art, Culture, and Technology Program
- Broad Institute of MIT and Harvard
- Center for Archaeological Materials
- Center for Bits and Atoms
- Center for Clinical and Translational Research
- Center for Collective Intelligence
- Center for Computational Science and Engineering
- Center for Constructive Communication
- Center for Energy and Environmental Policy Research
- Center for Environmental Health Sciences
- Center for Global Change Science
- Center for International Studies
- Center for Real Estate
- Center for Transportation & Logistics
- Concrete Sustainability Hub
- D-Lab
- Deshpande Center for Technological Innovation
- Division of Comparative Medicine
- Haystack Observatory
- Initiative on the Digital Economy
- Institute for Medical Engineering and Science
- Institute for Soldier Nanotechnologies
- Institute for Work and Employment Research
- Internet Policy Research Initiative
- Joint Program on the Science and Policy of Global Change
- Knight Science Journalism Program
- Koch Institute for Integrative Cancer Research
- Laboratory for Financial Engineering
- Laboratory for Information and Decision Systems
- Laboratory for Manufacturing and Productivity
- Laboratory for Nuclear Science
- Legatum Center for Development and Entrepreneurship
- Lincoln Laboratory
- Martin Trust Center for MIT Entrepreneurship
- Materials Research Laboratory
- McGovern Institute for Brain Research
- Microsystems Technology Laboratories
- MIT Center for Art, Science & Technology
- MIT Energy Initiative
- MIT Environmental Solutions Initiative
- MIT Kavli Institute for Astrophysics and Space Research
- MIT Media Lab
- MIT Office of Innovation
- MIT Open Learning
- MIT Portugal Program
- MIT Professional Education
- MIT Sea Grant College Program
- Nuclear Reactor Laboratory
- Operations Research Center
- Picower Institute for Learning and Memory
- Plasma Science and Fusion Center
- Research Laboratory of Electronics
- Simons Center for the Social Brain
- Singapore-MIT Alliance for Research and Technology Centre
- Sociotechnical Systems Research Center
- Whitehead Institute for Biomedical Research
- Women's and Gender Studies Program
- Architecture (Course 4)
- Art and Design (Course 4-B)
- Art, Culture, and Technology (SM)
- Media Arts and Sciences
- Planning (Course 11)
- Urban Science and Planning with Computer Science (Course 11-6)
- Aerospace Engineering (Course 16)
- Engineering (Course 16-ENG)
- Biological Engineering (Course 20)
- Chemical Engineering (Course 10)
- Chemical-Biological Engineering (Course 10-B)
- Chemical Engineering (Course 10-C)
- Engineering (Course 10-ENG)
- Engineering (Course 1-ENG)
- Electrical Engineering and Computer Science (Course 6-2)
- Electrical Science and Engineering (Course 6-1)
- Computation and Cognition (Course 6-9)
- Computer Science and Engineering (Course 6-3)
- Computer Science and Molecular Biology (Course 6-7)
- Electrical Engineering and Computer Science (MEng)
- Computer Science and Molecular Biology (MEng)
- Health Sciences and Technology
- Archaeology and Materials (Course 3-C)
- Materials Science and Engineering (Course 3)
- Materials Science and Engineering (Course 3-A)
- Materials Science and Engineering (PhD)
- Mechanical Engineering (Course 2)
- Mechanical and Ocean Engineering (Course 2-OE)
- Engineering (Course 2-A)
- Nuclear Science and Engineering (Course 22)
- Engineering (Course 22-ENG)
- Anthropology (Course 21A)
- Comparative Media Studies (CMS)
- Writing (Course 21W)
- Economics (Course 14-1)
- Mathematical Economics (Course 14-2)
- Data, Economics, and Design of Policy (MASc)
- Economics (PhD)
- Global Studies and Languages (Course 21G)
- History (Course 21H)
- Linguistics and Philosophy (Course 24-2)
- Philosophy (Course 24-1)
- Linguistics (SM)
- Literature (Course 21L)
- Music (Course 21M-1)
- Theater Arts (Course 21M-2)
- Political Science (Course 17)
- Science, Technology, and Society/Second Major (STS)
- Business Analytics (Course 15-2)
- Finance (Course 15-3)
- Management (Course 15-1)
- Biology (Course 7)
- Chemistry and Biology (Course 5-7)
- Brain and Cognitive Sciences (Course 9)
- Chemistry (Course 5)
- Earth, Atmospheric and Planetary Sciences (Course 12)
- Mathematics (Course 18)
- Mathematics with Computer Science (Course 18-C)
- Physics (Course 8)
- Department of Electrical Engineering and Computer Science
- Institute for Data, Systems, and Society
- Chemistry and Biology
- Climate System Science and Engineering
- Computation and Cognition
- Computer Science and Molecular Biology
- Computer Science, Economics, and Data Science
- Humanities and Engineering
- Humanities and Science
- Urban Science and Planning with Computer Science
- African and African Diaspora Studies
- American Studies
- Ancient and Medieval Studies
- Applied International Studies
- Asian and Asian Diaspora Studies
- Biomedical Engineering
- Energy Studies
- Entrepreneurship and Innovation
- Environment and Sustainability
- Latin American and Latino/a Studies
- Middle Eastern Studies
- Polymers and Soft Matter
- Public Policy
- Russian and Eurasian Studies
- Statistics and Data Science
- Women's and Gender Studies
- Advanced Urbanism
- Computational and Systems Biology
- Computational Science and Engineering
- Design and Management (IDM & SDM)
- Joint Program with Woods Hole Oceanographic Institution
- Leaders for Global Operations
- Microbiology
- Music Technology and Computation
- Operations Research
- Real Estate Development
- Social and Engineering Systems
- Supply Chain Management
- Technology and Policy
- Transportation
- School of Architecture and Planning
- School of Engineering
- Aeronautics and Astronautics Fields (PhD)
- Artificial Intelligence and Decision Making (Course 6-4)
- Biological Engineering (PhD)
- Nuclear Science and Engineering (PhD)
- School of Humanities, Arts, and Social Sciences
- Humanities (Course 21)
- Humanities and Engineering (Course 21E)
- Humanities and Science (Course 21S)
- Sloan School of Management
- School of Science
- Brain and Cognitive Sciences (PhD)
- Earth, Atmospheric and Planetary Sciences Fields (PhD)
- Interdisciplinary Programs (SB)
- Climate System Science and Engineering (Course 1-12)
- Computer Science, Economics, and Data Science (Course 6-14)
- Interdisciplinary Programs (Graduate)
- Computation and Cognition (MEng)
- Computational Science and Engineering (SM)
- Computational Science and Engineering (PhD)
- Computer Science, Economics, and Data Science (MEng)
- Leaders for Global Operations (MBA/SM and SM)
- Music Technology and Computation (SM and MASc)
- Real Estate Development (SM)
- Statistics (PhD)
- Supply Chain Management (MEng and MASc)
- Technology and Policy (SM)
- Transportation (SM)
- Aeronautics and Astronautics (Course 16)
- Aerospace Studies (AS)
- Civil and Environmental Engineering (Course 1)
- Comparative Media Studies / Writing (CMS)
- Comparative Media Studies / Writing (Course 21W)
- Computational and Systems Biology (CSB)
- Computational Science and Engineering (CSE)
- Concourse (CC)
- Data, Systems, and Society (IDS)
- Earth, Atmospheric, and Planetary Sciences (Course 12)
- Economics (Course 14)
- Edgerton Center (EC)
- Electrical Engineering and Computer Science (Course 6)
- Engineering Management (EM)
- Experimental Study Group (ES)
- Global Languages (Course 21G)
- Health Sciences and Technology (HST)
- Linguistics and Philosophy (Course 24)
- Management (Course 15)
- Media Arts and Sciences (MAS)
- Military Science (MS)
- Music and Theater Arts (Course 21M)
- Naval Science (NS)
- Science, Technology, and Society (STS)
- Special Programs
- Supply Chain Management (SCM)
- Urban Studies and Planning (Course 11)
- Women's and Gender Studies (WGS)
The Computer Science and Artificial Intelligence Laboratory (CSAIL) pursues fundamental research across the entire breadth of computer science and artificial intelligence. CSAIL is committed to leading the field both in new theoretical approaches and in the creation of applications that have broad societal impact.
CSAIL's current research activities span three principal areas:
- Artificial Intelligence (AI). This area of research aims to understand and develop systems—living and artificial—capable of intelligent reasoning, perception, and behavior. Specific research includes core AI computational biology, computer graphics, computer vision, human language technology, machine learning, medical informatics, robotics, and the semantic web.
- Systems. This area of research aims to discover common principles, models, metrics, and tools of computer systems, both hardware and software. Specific research includes compilers, computer architecture and chip design, operating systems, programming languages, and computer networks.
- Theory. This area of research studies the mathematics of computation and its consequences. Specific research includes algorithms, complexity theory, computations geometry, cryptography, distrusted computing, information security, and quantum computing.
CSAIL encourages student participation in its research projects. Undergraduates may become involved through the Undergraduate Research Opportunities Program (UROP) , and research assistantships are available to graduate students. CSAIL graduate students are typically enrolled in the departments of Electrical Engineering and Computer Science, Mathematics, Aeronautics and Astronautics, Brain and Cognitive Sciences, and Mechanical Engineering, and the MIT-Harvard Health Sciences and Technology Program.
Other related opportunities include:
- MIT App Inventor , a visual programming environment aimed at allowing new coders to build apps for smartphones and tablets
- The Internet Policy Research Initiative , an effort to conduct research and engage with public policy leaders on key issues in cybersecurity and technology

Print this page.
The PDF includes all information on this page and its related tabs. Subject (course) information includes any changes approved for the current academic year.

- Values of Inclusion
- 2020 Antiracism Task Force
- 2022 DEI Report
- Research News
- Department Life
- Listed by Recipient
- Listed by Category
- Oral History of Cornell CS
- CS 40th Anniversary Booklet
- ABC Book for Computer Science at Cornell by David Gries
- Books by Author
- Books Chronologically
- The 60's
- The 70's
- The 80's
- The 90's
- The 00's
- The 2010's
- Faculty Positions: Ithaca
- Faculty Positions: New York City
- Lecturer Position: Ithaca
- Post-doc Position: Ithaca
- Staff/Technical Positions
- Ugrad Course Staff
- Ithaca Info
- Internal info
- Graduation Information
- Cornell Learning Machines Seminar
- Student Colloquium
- Spring 2024 Colloquium
- Conway-Walker Lecture Series
- Salton 2023 Lecture Series
- Spring 2024 Artificial Intelligence Seminar
- Spring 2024 Robotics Seminar
- Spring 2024 Theory Seminar
- Big Red Hacks
- Cornell University - High School Programming Contests 2024
- Game Design Initiative
- CSMore: The Rising Sophomore Summer Program in Computer Science
- Explore CS Research
- ACSU Research Night
- Cornell Junior Theorists' Workshop 2023
- Researchers
- Ph.D. Students
- M.Eng. Students
- M.S. Students
- Ph.D. Alumni
- M.S. Alumni
- List of Courses
- Course and Room Roster
- CS Advanced Standing Exam
- Architecture
Artificial Intelligence
Computational biology, database systems, human interaction, machine learning, natural language processing, programming languages, scientific computing, software engineering, systems and networking, theory of computing.
- Contact Academic Advisor
- Your First CS Course
- Technical Electives
- CS with Other Majors/Areas
- Transfer Credits
- CS Honors Program
- CPT for International CS Undergrads
- Graduation Requirements
- Useful Forms
- Becoming a CS Major
- Requirements
- Game Design Minor
- Co-op Program
- Cornell Bowers CIS Undergraduate Research Experience (BURE)
- Independent Research (CS 4999)
- Student Groups
- UGrad Events
- Undergraduate Learning Center
- UGrad Course Staff Info
- The Review Process
- Early M.Eng Credit Approval
- Financial Aid
- Prerequisites
- The Application Process
- The Project
- Pre-approved Electives
- Degree Requirements
- The Course Enrollment Process
- Advising Tips
- Entrepreneurship
- Cornell Tech Programs
- Professional Development
- Contact MEng Office
- Career Success
- Applicant FAQ
- Computer Science Graduate Office Hours
- Exam Scheduling Guidelines
- Graduate TA Handbook
- MS Degree Checklist
- MS Student Financial Support
- Special Committee Selection
- Diversity and Inclusion
- Contact MS Office
- Ph.D. Applicant FAQ
- Graduate Housing
- Non-Degree Application Guidelines
- Ph. D. Visit Day
- Business Card Policy
- Cornell Tech
- Curricular Practical Training
- Fellowship Opportunities
- Field of Computer Science Ph.D. Student Handbook
- Field A Exam Summary Form
- Graduate School Forms
- Instructor / TA Application
- Ph.D. Requirements
- Ph.D. Student Financial Support
- Travel Funding Opportunities
- Travel Reimbursement Guide
- The Outside Minor Requirement
- CS Graduate Minor
- Outreach Opportunities
- Parental Accommodation Policy
- Special Masters
- Student Spotlights
- Contact PhD Office
Search form
You are here
The computing and information revolution is transforming society. Cornell Computer Science is a leader in this transformation, producing cutting-edge research in many important areas. The excellence of Cornell faculty and students, and their drive to discover and collaborate, ensure our leadership will continue to grow.
The contributions of Cornell Computer Science to research and education are widely recognized, as shown by two Turing Awards, two Von Neumann medals, two MacArthur "genius" awards, and dozens of NSF Career awards our faculty have received, among numerous other signs of success and influence.
To explore current computer science research at Cornell, follow links at the left or below.
Research Areas
Knowledge representation, machine learning, NLP and IR, reasoning, robotics, search, vision
Statistical genetics, sequence analysis, structure analysis, genome assembly, protein classification, gene networks, molecular dynamics
Computer Architecture & VLSI
Processor architecture, networking, asynchronous VLSI, distributed computing
Database systems, data-driven games, learning for database systems, voice interfaces, computational fact checking, data mining
Interactive rendering, global illumination, measurement, simulation, sound, perception
HCI, interface design, computational social science, education, computing and society
Artificial intelligence, algorithms
Programming language design and implementation, optimizing compilers, type theory, formal verification
Perception, control, learning, aerial robots, bio-inspired robots, household robots
Numerical analysis, computational geometry, physically based animation
Secure systems, secure network services, language-based security, mobile code, privacy, policies, verifiable systems
The software engineering group at Cornell is interested in all aspects of research for helping developers produce high quality software.
Operating systems, distributed computing, networking, and security
The theory of computing is the study of efficient computation, models of computational processes, and their limits.
Computer vision

espace - Curtin’s institutional repository
- espace Home
- Curtin Theses
A study of the effectiveness of computer laboratory classes as learning environments.
Access status.
This study focuses on the computer laboratory class as a learning environment in university courses. It involved the development and validation of two instruments, the Computer Laboratory Environment Inventory (CLEI) and the Attitude towards Computing and Computing Courses Questionnaire (ACCC). The CLEI has five scales for measuring students' perceptions of aspects of their laboratory environment. These are Student Cohesiveness, Open-Endedness, Integration, Technology Adequacy and Laboratory Availability. The ACCC has four scales, Anxiety, Enjoyment, Usefulness of Computers and Usefulness of the Course. The instruments were administered at three universities, one in Australia, one in England and one in the United States. The classes surveyed included those in which the development of software was the focus of study, such as Information Systems and Computer Science, and others in which the computer was used as a tool. With the exception of Laboratory Availability, all the environment variables were found to correlate significantly with all attitudinal variables. The only environment variable with significant association with achievement was Student Cohesiveness. However, the results showed that there were significant associations between the attitudinal variables, Anxiety, Enjoyment and Usefulness of the Course and achievement. Regression analysis supported the findings that the environment variables made a significant contribution to the attitudinal variables, and these in turn made a significant contribution to achievement. Further analysis using structural equation modelling suggests that computer laboratory environment affects achievement indirectly by directly affecting students' attitudes towards computers but even more so their attitude towards the course.The significance of this study is, that it is one of the first that has investigated the effectiveness of computer laboratory classes in a university setting in which the computer is central to the discipline being studied. The results demonstrate the importance of the laboratory environment in those courses in which the computer plays a major role. The CLEI will prove useful in the design and implementation of the laboratory component of a course and in the formative evaluation of such a course.
Related items
Showing items related by title, author, creator and subject.
- An interpretive study of the factors affecting the computer literacy of secondary school students. Newhouse, Christopher P. ( 1987 ) This study used interpretive research techniques to investigate the factors which affect the computer literacy of secondary students. The necessity that students to be prepared for life and work in a computer technology ...
- Evaluation of anthropometry activities for high school science: student outcomes and classroom environment Lightburn, Millard E. ( 2002 ) The study involved the evaluation of anthropometric activities for high school science. The activities actively engaged students in the process of gathering, processing and analyzing data derived from human body measurements, ...
- Perceptions of the learning environment, attitudes towards science, and understandings of the nature of science among prospective elementary teachers in an innovative science course Martin-Dunlop, Catherine S. ( 2004 ) The major purpose of this study was to evaluate the impact of a science course for prospective elementary teachers on their perceptions of the learning environment, attitudes towards science, and understandings of the ...
Show Statistical Information
- Menu Close
- Search
Bridging disciplines and accelerating discoveries in computer science.

Computer science research at the Johns Hopkins University is advancing computing technology, enabling new modes of thought, and transforming society. Our faculty conduct innovative, collaborative research aimed at solving large and complex interdisciplinary problems, drawing upon the university’s renowned strengths in areas including artificial intelligence, robotics, speech and language processing, medicine, and public health.
The department is rapidly growing, with current core research areas of theory and programming languages; systems and networking; computational biology and medicine; information security; natural language processing; machine learning, artificial intelligence, and data science; robotics, computer vision, and graphics; and human-computer interaction.
Researchers partners with colleagues in other engineering disciplines, as well as with investigators from the Johns Hopkins Krieger School of Arts and Sciences, the School of Medicine, and the Applied Physics Laboratory.
Cross-Departmental Centers and Institutes
- Center for Language and Speech Processing
- Laboratory for Computational Sensing and Robotics
- Johns Hopkins Information Security Institute
- Institute for Data Intensive Engineering and Science
- Malone Center for Engineering in Healthcare
- Human Language Technology Center of Excellence
- Center for Computational Biology
- Mathematical Institute for Data Science
Institute for Assured Autonomy
- Johns Hopkins Data Science and AI Institute
Research Areas
Theory & programming languages.
Focusing on the design, implementation, and use of computer programming languages.
Systems & Networkings
Our faculty are undertaking research into all aspects of computer systems and networks.
Computational Biology & Medicine
Faculty are engaged in a wide range of computational health and biology projects, from using data-driven tools to detect early signs of sepsis to DNA sequencing technology and evolutionary genomics.
Information Security
Hopkins researchers are working to safeguard our digital world.
Natural Language Processing
Creating innovative new technologies that will enable more natural interaction between human and computers.
Machine Learning, AI, & Data Science
Applying cutting-edge machine learning techniques to new datasets and domains.
Robotics, Vision, & Graphics
Research spans the areas of computer vision, computer graphics, and augmented and virtual reality.
Human-Computer Interaction
Placing people at the center of technological innovation.
Computer-Assisted Medicine
Our faculty are shaping the digital future across all aspects of health care.

The Institute for Assured Autonomy is operating in partnership with industry, government and academia to ensure the safe, secure, reliable, and predictable integration of autonomous systems into society by covering the full spectrum of research and application across the three pillars of technology, ecosystem, and policy & governance.
Login to your account
Change password, your password must have 8 characters or more and contain 3 of the following:.
- a lower case character,
- an upper case character,
- a special character
Password Changed Successfully
Your password has been changed
Create a new account
Can't sign in? Forgot your password?
Enter your email address below and we will send you the reset instructions
If the address matches an existing account you will receive an email with instructions to reset your password
Request Username
Can't sign in? Forgot your username?
Enter your email address below and we will send you your username
If the address matches an existing account you will receive an email with instructions to retrieve your username

- Institutional Access
Cookies Notification
Our site uses javascript to enchance its usability. you can disable your ad blocker or whitelist our website www.worldscientific.com to view the full content., select your blocker:, adblock plus instructions.
- Click the AdBlock Plus icon in the extension bar
- Click the blue power button
- Click refresh
Adblock Instructions
- Click the AdBlock icon
- Click "Don't run on pages on this site"
uBlock Origin Instructions
- Click on the uBlock Origin icon in the extension bar
- Click on the big, blue power button
- Refresh the web page
uBlock Instructions
- Click on the uBlock icon in the extension bar
Adguard Instructions
- Click on the Adguard icon in the extension bar
- Click on the toggle next to the "Protection on this website" text
Brave Instructions
- Click on the orange lion icon to the right of the address bar
- Click the toggle on the top right, shifting from "Up" to "Down
Adremover Instructions
- Click on the AdRemover icon in the extension bar
- Click the "Don’t run on pages on this domain" button
- Click "Exclude"
Adblock Genesis Instructions
- Click on the Adblock Genesis icon in the extension bar
- Click on the button that says "Whitelist Website"
Super Adblocker Instructions
- Click on the Super Adblocker icon in the extension bar
- Click on the "Don’t run on pages on this domain" button
- Click the "Exclude" button on the pop-up
Ultrablock Instructions
- Click on the UltraBlock icon in the extension bar
- Click on the "Disable UltraBlock for ‘domain name here’" button
Ad Aware Instructions
- Click on the AdAware icon in the extension bar
- Click on the large orange power button
Ghostery Instructions
- Click on the Ghostery icon in the extension bar
- Click on the "Trust Site" button
Firefox Tracking Protection Instructions
- Click on the shield icon on the left side of the address bar
- Click on the toggle that says "Enhanced Tracking protection is ON for this site"
Duck Duck Go Instructions
- Click on the DuckDuckGo icon in the extension bar
- Click on the toggle next to the words "Site Privacy Protection"
Privacy Badger Instructions
- Click on the Privacy Badger icon in the extension bar
- Click on the button that says "Disable Privacy Badger for this site"
Disconnect Instructions
- Click on the Disconnect icon in the extension bar
- Click the button that says "Whitelist Site"
Opera Instructions
- Click on the blue shield icon on the right side of the address bar
- Click the toggle next to "Ads are blocked on this site"
System Upgrade on Tue, May 28th, 2024 at 2am (EDT)
Computer laboratory environments: providing a suitable practical learning experience.
- Michael Newby
California State University, Fullerton, USA
Search for more papers by this author
The use of information and communications technologies in education has increased dramatically over the past decade. At the university level, computers are now used in most disciplines, either as an adjunct to the traditional lecture or to deliver the material on-line. Consequently, students are now required to master computer skills before they can master the subject being taught. To accommodate this need, tertiary institutions have provided access to computer laboratories. A computer laboratory is an expensive resource in terms of equipment and people, and should be used as effectively as possible. Computer laboratory classes may be organized as closed laboratories which are scheduled and staffed in the same way as other classes, or as open laboratories where students come and go as they please. This chapter reports the results of a study that investigated differences between students' perceptions of aspects of the learning environment of open and closed computer laboratories, and also the differences in student outcomes from courses that adopt these approaches. There was no significant difference in achievement between the two groups but there was a difference in their attitudes towards computers with those students gaining their practical experience from closed laboratories having a more positive attitude.
- Technology-supported learning environments in science classrooms in India Adit Gupta and Darrell Fisher 27 May 2012 | Learning Environments Research, Vol. 15, No. 2
- Online Viewing and Aesthetic Preferences of Generation Y and the Baby Boom Generation: Testing User Web Site Experience Through Eye Tracking Soussan Djamasbi, Marisa Siegel, Jeanine Skorinko and Tom Tullis 8 December 2014 | International Journal of Electronic Commerce, Vol. 15, No. 4
Recommended

Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- TECHNOLOGY FEATURE
- 28 June 2021
- Update 08 July 2021
Digital secrets of successful lab management
- Kendall Powell 0
Kendall Powell is a freelance writer in Boulder, Colorado.
You can also search for this author in PubMed Google Scholar
Illustration by The Project Twins
Christie Bahlai felt as if she was buried under a pile of virtual sticky notes. Like many group leaders, the computational ecologist appreciates that her team uses the messaging app Slack for virtual ‘water-cooler talk’. But she finds the app lacking when it comes to managing the various projects her laboratory is working on — threads, ideas and long-term goals get lost as conversations and memes rush on.
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
24,99 € / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
185,98 € per year
only 3,65 € per issue
Rent or buy this article
Prices vary by article type
Prices may be subject to local taxes which are calculated during checkout
Nature 595 , 138-139 (2021)
doi: https://doi.org/10.1038/d41586-021-01752-y
Updates & Corrections
Update 08 July 2021 : Shortly after this story was published, Bookkit was rebranded to have the same name as its parent company, Clustermarket. This has now been reflected in the text.
Related Articles

- Research management
- Communication
Science profits most when people of faith feel equally welcomed
Correspondence 11 JUN 24
Science and religion have profound differences — they should be kept apart

I was denied tenure — how do I cope?
Career Feature 06 JUN 24

Hybrid working from home improves retention without damaging performance
Article 12 JUN 24

Open access is working — but researchers in lower-income countries enjoy fewer benefits
Nature Index 11 JUN 24

How to track the economic impact of public investments in AI
Comment 10 JUN 24

‘Rainbow’, ‘like a cricket’: every bird in South Africa now has an isiZulu name
News 06 JUN 24

Misinformation poses a bigger threat to democracy than you might think
Comment 05 JUN 24
Misunderstanding the harms of online misinformation
Perspective 05 JUN 24
Postdoctoral positions at University of Pennsylvania
Postdoctoral positions funded by NIH are for study of meiosis and spermatogonial stem cells using mouse models. Generous stipend and benefits.
Philadelphia, Pennsylvania (US)
University of Pennsylvania - Department of Biomedical Sciences
Associate or Senior Editor, Nature Communications (Materials Science)
Associate or Senior Editor (Materials Science) Organization: Nature Communications Location(s): New York, Jersey City, Philadelphia, Beijing, Hong ...
New York (US)
Springer Nature Ltd
Associate or Senior Editor, Nature Medicine
Associate or Senior, Nature Medicine: Infectious Diseases New York, Shanghai or Beijing - Hybrid Working Application Deadline: August 1, 2024 Nat...
New York City, New York (US)
Post-doctoral Fellow
Funded investigator has an immediate opening for a post-doctoral fellow to develop therapies for genetic diseases.
Johns Hopkins School of Medicine, East Baltimore Campus
Johns Hopkins University Department of Medicine
Postdoctoral Fellow PhD
Houston, Texas (US)
Baylor College of Medicine (BCM)
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Academia.edu no longer supports Internet Explorer.
To browse Academia.edu and the wider internet faster and more securely, please take a few seconds to upgrade your browser .
Enter the email address you signed up with and we'll email you a reset link.
- We're Hiring!
- Help Center

"IMPROVING THE QUALITY OF COMPUTER LABORATORY FOR SENIOR HIGH SCHOOL STUDENTS OF MOUNT CARMEL SCHOOL OF MARIA AURORA, INC. (MCSMA): An Evaluation"

Dumpit, Sophia Lorainne V., Carrasco, Jedah Mae R. in Senior High School STEM Department (2019) Of Mount Carmel School of Maria Aurora, Inc. (MCSMA) Quantitative Research about the Improving the Quality of Computer Laboratory for Senior High School Students of Mount Carmel School of Maria Aurora, Inc. (MCSMA): An Evaluation. This quantitative research study was conducted to improve the quality of computer laboratory for Senior High School students of Mount Carmel School of Maria, Inc. (MCSMA). The aim of this study is to establish the development of computer laboratory to be use by the Senior High School students to help with their research paper and other study purposes. The study used the descriptive method to find some statistical data about their research study. Data were gathered through distribution of questionnaire to the selected senior high school students of Mount Carmel School of Maria Aurora, Inc. (MCSMA) and were statistically analyzed using the frequency and percentage mean. Total enumeration sampling was used in determining the sample population which was 151 Senior High School students. It resulted that the senior high students are favored in using the computer laboratory. The students also agree in developing the computer laboratory and also adding some equipment as they assume that it will be more convenient to comply the requirement inside the school. The students also willingness to add fees for the implementation of internet lab.
Related Papers
Larraine Sindac
This quantitative research paper was conducted to know the effect and the impact of having an internet, as a source of information, in Mount Carmel School of Maria Aurora, Inc.; to know if it would help the senior high school students do their requirements easily such as their research papers. The aim of this research is to ease the school works of these students. The result of this research will also help fulfill the lack of information in the library. In this way, the requirements of students are easier to conduct; the information needed is easier and faster to find inside the campus. Data were gathered through the random distribution of questionnaires to the Senior High Student of Mount Carmel School of Maria Aurora, Inc. (MCSMA) and were statistically analyzed through the use of Likert Scale. The results were gathered and were concluded that the internet is a tool that can ease the school works of the Senior High Student and was also a solution to fulfill the lack of information in the library.
Larraine Sindac , ephor vee
Galam, Von Angelo Z. and Sindac, Larraine V. in Senior High School STEM Department (2019) of Mount Carmel School of Maria Aurora, Inc. (MCSMA) Quantitative Research entitled A Study on the Implementation of Having Internet for the Requirements of the Senior High Students of Mount Carmel School of Maria Aurora (MCSMA), Inc. This quantitative research paper was conducted to know the effect and the impact of having an internet, as a source of information, in Mount Carmel School of Maria Aurora, Inc.; to know if it would help the senior high school students do their requirements easily such as their research papers. The aim of this research is to ease the school works of these students. The result of this research will also help fulfill the lack of information in the library. In this way, the requirements of students are easier to conduct; the information needed is easier and faster to find inside the campus. Data were gathered through the random distribution of questionnaires to the Senior High Student of Mount Carmel School of Maria Aurora, Inc. (MCSMA) and were statistically analyzed through the use of Likert Scale. The results were gathered and were concluded that the internet is a tool that can ease the school works of the Senior High Student and was also a solution to fulfill the lack of information in the library.
Justine Laroya
Justine Laroya , Kimberly Lazo
ABSTRACT Laroya, Justine Carl L., Dangco, Maria Isabelle D., and Kimberly Lazo in Senior High School, STEM Department (2019) of Mount Carmel School of Maria Aurora (MCSMA) Inc. Quantitative Research on the “The Effect of having not enough space to the Athletes of Mount Carmel School of Maria Aurora Inc. Sports is a widespread activity in the whole planet. There are plenty of events and games, and there`s also lot of human who involved with these games and wanted to be involved no matter what the situation is. Mount Carmel School of Maria Aurora Inc. is a school known for many positive things. One such thing is that the school promotes good quality of training through the right persons who supports the athletes. However, the space of the school isn`t enough for all the athletes. The study aimed to determine the Effects of having not enough space to the Athletes of Mount Carmel School of Maria Aurora Inc. Specifically, it sought to answer the demographic profile of the respondents in terms of name, sports that they use to play and if there is a significant effect of having not enough space to train for them. The descriptive method of research was used in this study. A descriptive method of research is a fact findings study with adequate and accurate interpretation of findings. It describes with emphasis that actually exist such as current conditions, situations or any phenomena. There were 50 selected respondents of this study. Studies find that having not enough space and physical facilities affects the performances of the athletes. Because the lack of space lessened their time to train and minimizes their limitation about giving their best. The study also finds that having not enough space affects the confidence of the athletes.
Research Paper
Zoe Vera Acain
ABSTRACT Acain, Zoe Vera S. and Calonge, Rav G. in Senior High School, STEM Department (2020) of Mount Carmel School of Maria Aurora (MCSMA). Quantitative Research on The Effect of the Voucher Program to the Household of the Students of Mount Carmel School of Maria Aurora (MCSMA), Inc. on their Pursuant to Senior High School Education. One of the leading problems in the education system of the Philippines is the large number of students that do not pursue higher education mainly due to the high tuition rate of their desired educational institutions. The Republic Act No. 10533 (RA 10533), also known as the Enhanced Basic Education Act of 2013, that extends the years of basic education from 10 years to 12 years has recently been implemented in the Philippines through the introduction of the senior high school. Critics of the law and affected families (especially those in the low-income bracket) argued the additional expenses. As a solution, the DepEd Order No. 11, series of 2015 (DO 11 s. 2015) introduced the Senior High School Voucher Program (SHS VP) to provide financial assistance to incoming senior high school students. The study aims to determine the effects of the program to the household, particularly the parents or guardians, of the students of Mount Carmel School of Maria Aurora (MCSMA), Inc. in their pursuance to senior high school. The study is also designed to determine if the SHS VP motivates students and their parents to pursue the additional two years of basic education, which is the senior high school. 2 To achieve the objectives of this research, a quantitative approach was used, with a questionnaire as the research tool. One hundred fiftyone questionnaires were administered randomly to students from different strand in Mount Carmel School of Maria Aurora (MCSMA), Inc. Results was represented through tables ang graph. The study concludes that most of the respondents are satisfied with the voucher program in terms of financial assistance. Since the program eased their financial burdens, it also widens the selection of the schools that a Filipino student can attend to. The financial assistance that are being provided by the government, such as the Voucher Program and the Free Tertiary Education, motivates and inspires the students to receive a higher education so that they can be more competitive in the employment market. Keywords: Senior High School Voucher Program (SHS VP), Tuition Fee, Senior High School
Lian Flores
Ellaine Obar
Obar, Ellaine Therese D. and Lucas, Mark John D. in Science, Technology, Engineering and Mathematics (STEM) strand of Mount Carmel School of Maria Aurora, Inc. Quantitative research entitled ‘The Positive Effect of Contemporary Teaching Methods in the Academic Performances of Senior High School Students in Mount Carmel School of Maria Aurora, Inc.’ This quantitative research study was conducted to determine if the Senior High School students of Mount Carmel School of Maria Aurora, Inc affect their academic performances in applying contemporary teaching methods. It involved the Grade 11 ad 12 students because they have lot of experiences towards the traditional teaching methods to modern teaching methods. The study focuses mainly on the problem encountered by every student about teaching methods. The data were gathered through questionnaire and surveying random students of Senior High School Department In the end of the result of the study is positive wherein contemporary teaching methods help the students in their academic performances. Keywords: contemporary, traditional,modern, teaching methods, experiences, academic performances
desiree tamayo
Tamayo, Desiree C. and Nisperos, Niña Kerstin M. in Science, Technology, Engineering, and Mathematics (STEM) strand of Mount Carmel School of Maria Aurora, Inc, (MCSMA) Quantitative research about The Positive Effect of Implementing Later School Start Time to Senior High School Students of Mount Carmel School of Maria Aurora, Inc. This quantitative research study was conducted to find out what are the effects of implementing later school start time to senior high school students of Mount Carmel School of Maria Aurora, Inc. The study aims to help the students get the sleep they need to function every day, the study also aims to help reduces the number of absent and late students in the school. The data were gathered through questionnaires and surveying random students of Senior High School Department. In the end the result of the study is positive wherein implementing later school start time would help the student to get the suggested sleep they need to function better every day, in addition it will also help them have a good health and away from depression and anxiety. Keyword: school, later, start time, students, sleep
Oscar Vallejo
Loading Preview
Sorry, preview is currently unavailable. You can download the paper by clicking the button above.
RELATED PAPERS
Kendall Jenner
Godfrey Nacino
Chika Odedo
George Fulford
Obasi Nnanna
MSc Educational Leadership and Management
Mark Camilleri
Terje Mølster
Joselyne Teh , Brixter Luquing
Eshetu Mandefro
Dalaguete PH
GEORGE LUMAYAG
Zongo Nebanat
Edward Penn-Timity
Nana Frempong Badasu
Dr. Muhammad Kristiawan, M.Pd.
ijedict.dec.uwi.edu
Stewart Marshall
Dr.Olukayode S O L O M O N Aboderin
thandazile precious
K. M. Lawal
lutenta munsaka
International Journal of
Frank Antwi Boasiakoh
… : sharing the learning …
Paula Wilcox
Inocencia M Canon
Faremi Seun Samuel
Edwin Tatel III
Carla Basili , Thomas Hapke , Sabina Cisek
International Journal of Science and Research Publications
Ayivi Charles
Ticha Ludmila , Carla Basili , Hana Landová , Josep Vives-Gràcia , Michaela Dombrovská , Sabina Cisek
IJAR Indexing
umoru titus
Mark Valentine Aikins
Mvelo Walaza
RELATED TOPICS
- We're Hiring!
- Help Center
- Find new research papers in:
- Health Sciences
- Earth Sciences
- Cognitive Science
- Mathematics
- Computer Science
- Academia ©2024
- Dean’s Office
- External Advisory Council
- Computing Council
- Extended Computing Council
- Undergraduate Advisory Group
- Break Through Tech AI
- Building 45 Event Space
- Infinite Mile Awards: Past Winners
- Frequently Asked Questions
- Undergraduate Programs
- Graduate Programs
- Educating Computing Bilinguals
- Online Learning
- Industry Programs
- AI Policy Briefs
- Envisioning the Future of Computing Prize
- SERC Symposium 2023
- SERC Case Studies
- SERC Scholars Program
- SERC Postdocs
- Common Ground Subjects
- For First-Year Students and Advisors
- For Instructors: About Common Ground Subjects
- Common Ground Award for Excellence in Teaching
- New & Incoming Faculty
- Faculty Resources
- Faculty Openings
- Search for: Search
- MIT Homepage

Labs & Centers

Computer Science and Artificial Intelligence Laboratory (CSAIL)

Laboratory for Information and Decision Systems (LIDS)
MIT Quest for Intelligence
- View MIT Quest

MIT-IBM Watson AI Lab
- View MIT-IBM Watson AI Lab

Abdul Latif Jameel Clinic for Machine Learning in Health (Jameel Clinic)
- View Jameel Clinic

Sociotechnical Systems Research Center (SSRC)
- View Center for Biomedical Innovation
Related Stories

School of Computer Science
College of computing.
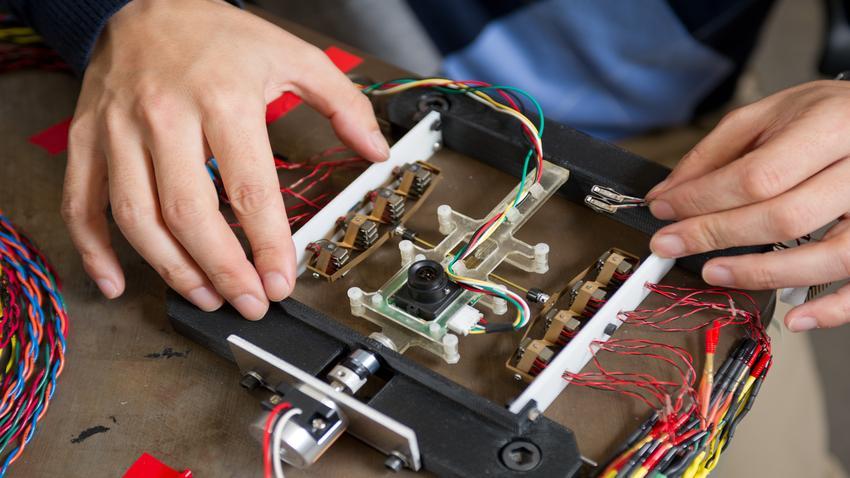

Groups & Labs
Comparch lab.
Faculty: Tom Conte, Hadi Esmaeilzadeh, Hyesoon Kim, Santosh Pande, Milos Prvulovic, Kishore Ramachandran
The Computer Architecture (comparch) Lab conducts research on all aspects of future microprocessor technology including performance, power, multi-threading, chip-multiprocessing, security, programmability, reliability, interaction with compilers and software, and the impact of future technologies.
Data Systems and Analytics Group
Faculty: Joy Alruraj, Xu Chu, Constantine Dovrolis, Vladimir Kolesnikov, Ling Liu, Kexin Rong, Shamkant Navathe, Calton Pu, Jun Xu
We live in an era of unprecedented access to data. Our group tackles new challenges in organizing and leveraging the vast amounts of information at our disposal using data management and machine learning techniques. We participate in a number of cross-disciplinary research efforts, and closely collaborate with other groups at Georgia Tech.
Distributed Data Intensive Systems Lab
Faculty: Ling Liu, Calton Pu
DiSL offers research expertise in distributed and Internet computing systems and distributed data intensive systems. Most of the research projects conducted in DiSL have strong emphasis on systems issues such as scalability, reliability, security, availability and efficiency, and data management issues such as data storage, data mining and data analysis. We are interested in theories and techniques that not only make the distributed systems scalable and efficient but also reliable and secure.
Efficient and Intelligent Computing (EIC) Lab
Faculty: Yingyan (Celine) Lin
The EIC lab in the School of Computer Science at Georgia Tech focuses on developing efficient machine learning (ML) techniques via cross-layer innovations, spanning from artificial intelligence (AI) algorithms to AI hardware accelerators and AI chip design , and aims to foster green AI and ubiquitous AI-powered intelligence.
Embedded Pervasive Lab (EPL)
Faculty: Kishore Ramachandran
Research in the Embedded Pervasive Lab (EPL) covers a range of topics including distributed programming idioms, networked embedded sensors, P2P video streaming, middleware for efficient large-scale stream processing, opportunistic networking, virtualization technologies, and smart storages incorporating Flash-based SSDs. EPL is producing systems for a world where machines, devices and networking technologies work in concert to help individuals use them with as little instruction as possible.
Hardware Security Lab
Faculty: Daniel Genkin
Research in secure hardware design, microarchitectural side-channel attacks, and applied cryptography, at Georgia Institute of Technology.
Internet Intelligence Research Lab
Faculty: Zachary Bischof, Alberto Dainotti, Cecilia Testart, Amanda Meng
The Internet Intelligence Research Lab focuses on understanding and improving the security and reliability of the Internet.
Korvo Research
Faculty: Greg Eisenhauer, Ada Gavriloska, Matthew Wolf, Jeff Young
The Korvo research group of researchers focuses on pushing the boundaries of computer systems research. We take an experimental approach to computer systems, working heavily with industry and governmental stakeholders with real-world problems as motivation for our efforts to refine and fundamentally restructure the way operating systems work. Our interests cover the areas of Cloud Computing, High Performance Computing, and Internet of Things, with particular focus on building systems that can deal with tight orchestration of large streams of data, wherever that data source may come from.
Systems Software & Security Lab
Faculty: Taesoo Kim
In the “SS Lab,” housed within the Georgia Tech Information Security Center, we build practical systems with focuses on security, performance, robustness, or often just for fun. Our research projects have been published in top academic conferences, and have made great impacts on real programs, such as Firefox and Android, that people use every day.
Networks Lab
Faculty: Mostafa Ammar, Constantine Dovrolis, Jim Xu, Ellen Zegura, Russ Clark
Our networks group aims to understand and advance the theory and practice of networks, whether they be mobile, wireless, Internet, campus, home, physical, virtual, human-constructed or naturally occurring. We combine modeling with measurement and experimentation to connect insights to novel approaches to solving problems. Our areas of expertise include network algorithmics, multicast, multimedia, disruption-tolerant networking, mobile computing, network operations, Internet economics, and network science.
Faculty: Jacob Abernethy, Merrick Furst, Zvi Galil, Richard Lipton, Will Perkins, Dana Randall, Mohit Singh, Sahil Singla, Prasad Tetali, Jan van den Brand, Santosh Vempala
Our theory group has expertise spanning foundations and applications of algorithms and computational complexity theory, including combinatorial optimization, approximation algorithms, randomized algorithms, stochastic processes, spectral methods, algorithmic game theory, high-dimensional geometry, network models, and cryptography. The theory group also constitutes some of the core faculty of both the College of Computing’s Algorithms & Randomness Center (ARC) and Georgia Tech’s Ph.D. program in Algorithms, Combinatorics and Optimization (ACO).
Laboratories
Current labs.
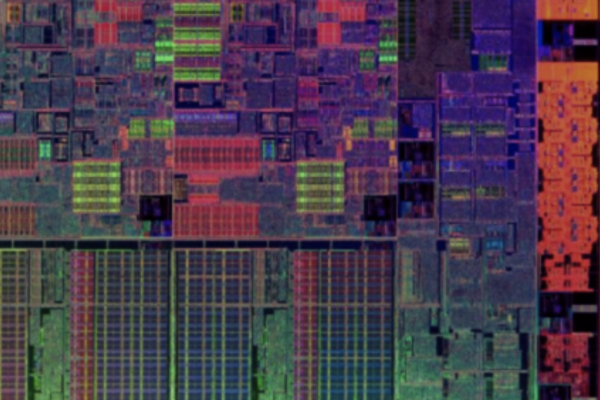

The Bionic Vision Lab is an interdisciplinary group that combines expertise in computational neuroscience, neuroengineering, and human-computer interaction. The lab’s particular focus is on bionic vision, in which brain-computer interfaces serve as a tool to study the neural mechanisms of visual perception in people with vision loss, with the ultimate goal of restoring useful vision to people who are blind.

Our expertise is in matrices, graphs, and computation. Our research is in combinatorial scientific computing, high-performance graph algorithms, tools and software for computational science and engineering, and numerical linear algebra. We are interested in applications across the entire spectrum of computational science and data analysis.

Focus on mathematical modeling and machine learning, applied to networked systems in biology and medicine. Current projects include development of algorithms and software for both simulation and inference of discrete stochastic systems, uses of machine learning applied to medical data to improve the care of trauma patients, mathematical modeling of brain processes in migraine, unraveling neural communication in learning, inference of network structure of communities of neurons, and mathematical modeling of the process of cell polarization.

Our research focuses on machine learning and data mining, social networks, brain networks, and bioinformatics.

UCSB's GameLab provides a supportive and educational environment for students to pursue their creative endeavors in game development and learn more about the development of games. We welcome people of all skills and skill levels who are interested in any aspect of game development, whether it be programming, art, writing, sound design, production, or the many other fields that go into crafting great games.

Advancing machine learning theory, algorithms and applications.

The MOMENT Lab focuses on two areas of research: mobile networking, and information and communication technologies for development (ICTD). In many projects, we apply our nearly two decades of expertise in mobile and wireless networking to solve problems in ICTD. Within this space, the MOMENT lab works on the development of network solutions, primarily but not exclusively wireless, suitable to the constraints of developing and underdeveloped regions of the world.

Investigating the design, analysis, and implementation of Programming Languages.

Research conducted includes web search, information retrieval and mining.

The Computer Security Group at UCSB works on tools and techniques for designing, building, and validating secure software systems. The group's research focus is on vulnerability analysis, cloud security, malware and the underground economy, smartphone and IoT security, and the security of web-based applications.

UCSB Systems and Networking Lab builds state-of-the-art computer systems, especially those focused on autonomous network management and privacy protection. The group's research interests are in computer systems and networking. A particular focus is on problems in systems and networking that intersect with machine learning and privacy. We are working on self-driving networks, which studies how machine learning can be applied to automate network-management tasks at scale.

Research at VLab focuses on automated verification techniques and their application to software in order to improve software security and dependability.

Computer Systems Lab @ Yale
A research laboratory spanning Computer Science and Electrical Engineering
Welcome to the Computer Systems Lab (CSL) at Yale University . CSL is an interdisciplinary laboratory with faculty from both Electrical Engineering and Computer Science that have a shared research interest in computer systems. CSL research encompasses both theory and practice in a wide range of domains including architecture, biologically-inspired computing, certified systems software, distributed systems, networking, operating systems, programming languages, security, storage, and VLSI.
Graduate students working in CSL are admitted to either the Electrical Engineering or Computer Science department, but are encouraged to apply to the department that is closest to their research interests. Most faculty members in CSL have appointments in both CS and EE, making it easy for students to work across traditional disciplinary boundaries.

Rotate Panorama
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- Healthcare (Basel)

Digital Management Systems in Academic Health Sciences Laboratories: A Scoping Review
Margareth timóteo.
1 Clinical Research Unit, Antônio Pedro Hospital, Fluminense Federal University, Niteroi 24020-140, Brazil; moc.liamg@oetomitagram (M.T.); rb.ffu.di@zorieuqdl (L.D.); moc.liamg@lraepecioj (J.d.S.); moc.liamg@4revilosnurb (B.O.); rb.ffu.di@jeloineb (B.O.)
2 Post-Graduation Program in Medical Sciences, Fluminense Federal University, Niteroi 24020-140, Brazil
Emanuelle Lourenço
3 Post-Graduation Program in Dentistry, Fluminense Federal University, Niteroi 24020-140, Brazil; rb.moc.oohay@tellets_elleuname
Ana Carolina Brochado
4 Post-Graduation Program in Science and Biotechnology, Fluminense Federal University, Niteroi 24020-140, Brazil; [email protected] (A.C.B.); moc.liamtoh@aacil_ataner (R.B.)
Luciana Domenico
Joice da silva, bruna oliveira, renata barbosa, pietro montemezzi.
5 Independent Researcher, 24128 Bergamo, Italy; [email protected]
Carlos Fernando de Almeida Barros Mourão
Gutemberg alves, associated data.
The data presented in this study are openly available in the Open Science Framework (OSF) database, at doi:10.17605/OSF.IO/KPC3Q.
Good laboratory practices (GLP) increase the quality and traceability of results in health sciences research. However, factors such as high staff turnover, insufficient resources, and a lack of training for managers may limit their implementation in research and academic laboratories. This Scoping Review aimed to identify digital tools for managing academic health sciences and experimental medicine laboratories and their relationship with good practices. Following the PRISMA-ScR 2018 criteria, a search strategy was conducted until April 2021 in the databases PUBMED, Web of Sciences, and Health Virtual Library. A critical appraisal of the selected references was conducted, followed by data charting. The search identified twenty-one eligible articles, mainly originated from high-income countries, describing the development and/or implementation of thirty-two electronic management systems. Most studies described software functionalities, while nine evaluated and discussed impacts on management, reporting both improvements in the workflow and system limitations during implementation. In general, the studies point to a contribution to different management issues related to GLP principles. In conclusion, this review identified evolving evidence that digital laboratory management systems may represent important tools in compliance with the principles of good practices in experimental medicine and health sciences research.
1. Introduction
Laboratory research plays an essential role in providing evidence for translational medicine and sustainable solutions to healthcare [ 1 ]. However, the reliance on experimental medicine demands increased traceability and data integrity, ensuring the quality of transferrable results to the clinical setting. In recent years, the scientific community experienced awareness regarding a reproducibility crisis related to factors such as the pressure for publication, low statistical power, and insufficient supervision [ 2 ]. On the other hand, adequate management, training, and good practices may increase data quality by improving workflow, avoiding errors, and providing traceability [ 2 ].
Good laboratory practices (GLP) may be defined as a quality system encompassing organizational processes and conditions under which studies are planned, executed, monitored, registered, and reported [ 3 ]. The Principles of Good Laboratory Practice were first developed by a group of GLP experts led by the USA, established in 1978 under the Special Program on the Control of Chemicals, based on the FDA’s regulations for non-clinical laboratory studies. The Organization for Economic Cooperation and Development (OECD) published the Principles of Good Laboratory Practice and Compliance Monitoring in January 1998 [ 3 ]. Since then, it represents the primary set of standards available worldwide to ensure quality, reliability, and integrity, providing a solid approach to the management of research laboratories [ 4 ].
However, academic laboratories experience several critical barriers to developing and implementing a GLP-compliant infrastructure [ 5 ]. These limitations include poor training on management, lack of funding for compliance costs, and high staff turnover due to a dependence on students as temporary personnel [ 6 ]. Therefore, laboratory managers at academic centers should explore tools that facilitate supervision and identify critical steps in the laboratory workflow. In this context, digital systems are among the most important tools available for efficient management, ranging from dedicated computer programs to smartphone applications. Laboratory information management systems (LIMS) offer databases and automation [ 7 ] that allow experimental data tracking and storage [ 8 ]. Other software and digital services that fall outside of the original LIMS classification provide a broader offer of solutions to laboratory management [ 6 , 9 ], coping with other aspects of quality assurance related to communication, staff, multiuser equipment schedule and maintenance, standard procedures, and inventory control, which are fundamental in the full spectrum of a laboratory’s workflow [ 10 , 11 ].
Despite the potential effectiveness of these digital tools in meeting specific aspects of laboratory management, it remains unclear how these systems may directly or indirectly contribute to adherence to the GLP principles. In this context, the present review aimed to provide evidence on the theme by scoping the scientific literature for the available digital tools designed to manage health sciences and experimental medicine laboratories and discuss the assessments of effectiveness, acceptance, and their potential for compliance to different aspects of good laboratory practices.
2. Materials and Methods
2.1. protocol and registration.
This review followed the PRISMA recommendations for scoping reviews (PRISMA-ScR) [ 12 ], as shown in the Supplementary Table S1 . The study protocol was registered in the Open Science Framework database under the Digital Object Identifier doi:10.17605/OSF.IO/KPC3Q on 15 July 2020.
2.2. Sources of Information and Research Strategy
The broad question that guided the review was: “Are there available digital tools for the management of academic health sciences laboratories?” Strategies were developed to search for data sources in three different databases: PUBMED ( www.ncbi.nlm.nih.gov/pubmed (accessed on 26 April 2021)), Web of Science (WoS) (clarivate.com/webofsciencegroup (accessed on 26 April 2021)), and the Virtual Health Library (VHL) (bvsalud.org (accessed on 26 April 2021)). The research was carried until April 2021. Grey literature was consulted through the OpenGrey Database (available at http://www.opengrey.eu/ (accessed on 24 May 2021)). The search keys are described in Table 1 , with various combinations of Medical Subject Headings (MeSH) descriptors selected to cover as many articles as possible coping with management software approaches in academic or research settings.
Search keys applied to the three consulted databases.
| Database | Search Key |
|---|---|
| PUBMED | (laborator*[tiab] OR Laboratories[mh]) AND (management[tiab] OR “Organization and Administration”[mh] OR “Information Management”[mh]) AND (software[tiab] OR computer*[tiab] OR virtual[tiab] OR Software[mh] OR “Mobile Applications”[mh]) AND (academic OR Universities[mh] OR research[tiab] OR research[mh] OR “Biomedical Research”[mh] OR “Translational Medical Research”[mh]) AND (health OR clinic*) |
| Web of Science | TOPIC: ((laboratory) AND (management OR “Organization and Administration” OR “Information Management”) AND (software OR computer OR virtual OR “Mobile Applications”) AND (academic OR University OR research OR “Biomedical Research” OR “Translational Medical Research”) AND (health OR clinic)). Time stipulated: all years. Indices: SCI-EXPANDED, SSCI, A&HCI, CPCI-S, CPCI-SSH, ESCI. |
| Virtual Health Library | (laboratory) AND (management OR organization) AND (software OR computer OR virtual OR “Mobile Applications”) AND (academic OR University OR research) AND (health OR clinic) |
2.3. Selection of Sources of Evidence
The eligibility criteria were determined on a PIO (Population, Intervention, Outcome) variant of the PICO framework for the selection of studies, more adequate for qualitative reviews [ 13 ].
A structured question was produced, in which, Population (P): academic health sciences laboratories, Intervention (I): the use of digital tools, and Outcomes (O): management for quality. After the references were retrieved from the database search, a group of five trained and calibrated reviewers read all titles and abstracts, applying the eligibility criteria, which included complete works on digital tools that aid in the administration of laboratories in academic or research environments, in health or biomedical sciences, including collections and biorepositories. Studies were excluded if they (i) were entirely out of the subject, (ii) did not address laboratory management, (iii) did not deal with software or digital tools, and (iv) were not proposed or discussed for health sciences or biomedical research. Additionally, articles on software that exclusively assessed experimental data management were considered outside the scope of this review. The inter-examiner reliability was assessed through simultaneous assessment of references by five evaluators, obtaining a Cohen’s Kappa coefficient of 0.93. Doubts and disagreements were resolved in weekly meetings conducted during this stage.
2.4. Critical Appraisal
A critical appraisal was conducted with the selected references, applying an instrument described by Whittemore and Knafl [ 14 ], considering two relevant criteria: (i) methodological and theoretical soundness and (ii) relevance of the data to the proposed question of the review. The methodological assessment considered whether studies presented adequate identification and traceability of the software, evaluating effectiveness, applicability, or acceptance. The adherence to the review’s question was considered according to the description of management functions, target users and environment, and software limitations. Each present parameter was scored with 1 point, to a maximum of 4 points. No study was excluded based on this assessment classification, even though the score was included as a variable in the data analysis stage. In general, studies of lower scores contributed less to the analytical process.
2.5. Synthesis of Results and Data Charting
The main characteristics of the selected studies were collected and tabulated, including year and country of conduction, name and type of digital tool, topics of laboratory management issued by the software, target public and environment of application, accessibility, and whether the software was free or paid. The data extraction was performed in conjunction with five authors in regular meetings. A specific table was produced solely with the studies that performed evaluations of effectiveness or acceptance, with the respective outcomes. A chart was produced connecting the management topics issued by the different tools and the respective sections/chapters from the Organization for Economic Cooperation and Development (OECD) GLP Principles [ 3 ].
Figure 1 shows the results for the search strategy and screening of databases. The PUBMED database provided 855 entries, while 183 entries were identified in WoS and 550 in the VHL. After combining the 3 results, 352 duplicate articles were identified and excluded. After applying the exclusion criteria, 523 articles were considered off-topic, appearing in searches because of common words and often dealing with clinical/hospital-related issues, but not with experimental medicine. Of the total articles identified, 160 were excluded because they did not deal with software or digital systems, and 534 did not speak about management. From the screening result, 19 articles were selected to compose this Scoping Review, and 2 additional articles were identified manually upon reading the selected references. The twenty-one elected references included studies proposing new software or revisiting already available tools for novel management applications. Some authors also evaluated the impact of changes during and after implementing the systems, either qualitatively or quantitatively.

PRISMA flowchart of study screening and selection.
Table 2 shows a critical appraisal performed for the selected articles at the methodological level and relevance to our broad question. Of the 21 articles selected, 9 evaluated the effectiveness and pointed out the limitations. Another eight did not evaluate but described limitations, and four studies did not evaluate or point out the limitations of the systems used, only describing the implementation or development of the systems in an expository manner. Nevertheless, all articles adequately identified the investigated software, their management purposes, and the environment/professionals served by its functionalities were considered relevant and contributed to some extent to the qualitative discussion on the theme.
Critical appraisal of the sources of evidence.
| Adequacy to the Research Question | Methodological Soundness | ||||
|---|---|---|---|---|---|
| Reference | Description of Software Limitations | Description of Functions and Users/Environment | Evaluation of Applicability, or Acceptance | Adequate Identification and Traceability | Final Score |
| Delorme and Cournoyer [ ] | 1 | 1 | 1 | 1 | 4 |
| Godmann et al. [ ] | 1 | 1 | 0 | 1 | 3 |
| Nayler and Stamm [ ] | 1 | 1 | 0 | 1 | 3 |
| Selznick et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Anderson et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Viksna et al. [ ] | 0 | 1 | 0 | 1 | 2 |
| Milisavljevic et al. [ ] | 1 | 1 | 0 | 1 | 3 |
| Yousef et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Machina and Wild [ ] | 1 | 1 | 0 | 1 | 3 |
| Allwood et al. [ ] | 1 | 1 | 0 | 1 | 3 |
| Calabria et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Perkel [ ] | 0 | 1 | 0 | 1 | 2 |
| Boutin et al. [ ] | 0 | 1 | 0 | 1 | 2 |
| Catena et al. [ ] | 1 | 1 | 0 | 1 | 3 |
| Dirnagl et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Manca et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Paul et al. [ ] | 0 | 1 | 0 | 1 | 2 |
| Gaffney et al. [ ] | 1 | 1 | 0 | 1 | 3 |
| Dennert Friedrich and Kumar [ ] | 1 | 1 | 1 | 1 | 4 |
| Timoteo et al. [ ] | 1 | 1 | 1 | 1 | 4 |
| Cooper et al. [ ] | 1 | 1 | 0 | 1 | 3 |
To quantify the criteria, “1” means present, and “0” means absent.
Table 3 describes the main characteristics of the twenty-one selected studies related to the present research question. It can be observed that the selection ranged from studies of the earlier days of the use of personal computers in laboratories [ 15 , 16 ] to current cloud computing and mobile applications [ 25 , 29 ]. In addition, some references studied the complexities of the concomitant use of several integrated tools [ 11 , 30 ]. In accordance with the search criteria, the studied environments consisted of academic, health-related laboratories, as well as biorepositories and biobanks. Consistently, the target users were managers and staff common to these laboratories, including technicians, researchers, doctors, and students.
Main characteristics of the selected studies.
| Reference | Country | Software | Availability | Managed Activity | Environment | Target Users | Costs |
|---|---|---|---|---|---|---|---|
| Delorme and Cournoyer [ ] | UK | Customer Information Control System/Virtual Storage (CICS/VS) | Desktop | Tax and administrative tasks, quality control of data and techniques, epidemiological assistance, and teaching and research in the different subspecialties of microbiology. | Microbiology laboratory at a university hospital | Medical Doctors, researchers, and students | Charged |
| Godmann et al. [ ] | USA | LabFlow | Desktop | Workflow in large-scale biology research laboratories. | Research Laboratory | Researchers and laboratory users | Free |
| Nayler and Stamm [ ] | Germany | ScienceLab Database (SLD) | Desktop | Stock of reagents and biological samples, protocols, library, vendor information. | Molecular biology laboratory | Laboratory professionals | Charged |
| Selznick et al. [ ] | USA | Cell Culture Laboratory Management System (CCLMS) | Desktop | Cell culture laboratory management: modules for registering cell counts, frozen cell records, user records, and culture vessel specifications. | Cell culture laboratory | Researchers and users of cell culture laboratories | Custom prototype |
| Anderson et al. [ ] | USA | MGEA | Desktop | Experimental workflow, integration with laboratory equipment, storage, and statistical analysis of experimental data. | Genetic research laboratory | Researchers, laboratory professionals, biostatistics, students. | Charged |
| Viksna et al. [ ] | UK | Patient and Sample System for Information Management (PASSIM) | Desktop/ online | Study participants, samples, and results. | Biorepository and biomedical research labs | Researchers and students | Free and open source |
| Milisavljevic et al. [ ] | Canada | Laboratory Animal Management Assistant (LAMA) | Online | Management of mouse colonies. | Biotery. | Researchers | Free |
| Allwood et al. [ ] | Canada | Lennie | Smartphone | Maintenance and management of animal colonies. | Vivarium. | Researchers | Free |
| Yousef et al. [ ] | USA | LINA | Desktop | Track collections of biologically relevant materials. | Molecular biology academic laboratories. | Medical Doctors, researchers and students | Free |
| Machina and Wild [ ] | USA | Electronic Laboratory Notebook (ELN) | Desktop | Automation of lab tests; register of equipment-related data (use, and calibration). Laboratory inventories. | General laboratories. | Researchers and laboratory users | Charged |
| Calabria et al. [ ] | USA | AdLIMS | online | Biological samples; metadata from patient samples; experimental procedures, workflow, and data for DNA samples. | Genetic sequencing laboratories. | Researchers and users of cell culture laboratories | Charged |
| Perkel [ ] | USA | Quartzy; LabGuru; LINA; StrainControl; CISPro; mLIMS; OpenFreezer | Online/ smartphone | Sample tracking and inventory. | Research laboratories and Academic Institutions | All levels of laboratory staff | Free and charged tools |
| Boutin et al. [ ] | USA | STARLIMS; GIGPAD; Crimson; Constrack; EMSI; Biobank portal | Online/ smartphone | Genomic data transfer, sequencing, genotyping, sample inventory, workflow, DNA and RNA sample processing and tracking; patient data. | Research laboratories, biobanks, collection clinics, hospitals | Coordinators, research subjects, researchers, IT staff. | A combination of custom/ free/ charged tools |
| Catena et al. [ ] | Switzerland | AirLab | Online/ smartphone | Reagent and sample inventory; database of antibodies. | Research laboratories, mainly molecular. | Researchers and laboratory students | Free |
| Dirnagl et al. [ ] | Germany | LabCIRS (Laboratory critical incident report) | Online | Risk/error management. | Research Laboratories | Research groups, laboratories, and institutions | Free |
| Manca et al. [ ] | USA | Laboratory Center (LC) | Online | Virtual biorepository. | Antibacterial Research Laboratory; biorepository | Researchers | Charged |
| Paul et al. [ ] | USA | BlazeLIMS; FreezerPro; WebLIMS; BioTracer | Cloud | Biobanking. | Biobanks | Doctors and researchers | Charged |
| Gaffney et al. [ ] | USA | GEM-NET | Online | Access control, data and protocol storage, project monitoring, teamwork, internal communication, engagement, and biorepositories. | Academic Research Laboratories—Biorepository of specimens | Researchers and laboratory students | A combination of free/charged tools |
| Timoteo et al. [ ] | Brazil | Quartzy | Online | Staff and workflow management of an academic research lab including documentation, equipment, inventory, and communication. | Academic Research Laboratories | Researchers, and laboratory users. | Free/charged versions |
| Dennert, Friedrich and Kumar [ ] | USA | Database—SDLC System | Online; LAN. | Inventory Management System | Research laboratories; Biorepository of specimens | Researchers and laboratory staff | Charged |
| Cooper et al. [ ] | USA | LabDB | Online; LAN. | Manages experimental data and organizes personnel and inventory. | Research laboratories. | Researchers | Charged |
Thirty-three programs/systems were identified in the twenty-one studies, with eight exclusively available for installation on desktop computers and the rest available online, including cloud-based systems, that is, with storage on online servers and availability on demand. Twenty-one of the studied systems were commercially available, charged programs/services, while twelve were free-of-charge for some of their functionalities. Among the non-charged software, two were custom systems designed exclusively for the studied laboratory (Biobank Portal and CCLMS).
Table 4 summarizes the results of the nine studies that assessed the impact of implementing computerized management systems. All of them reported positive results with the use of digital-assisted management. However, problems were identified related to technical constraints (either hardware or software) and limited acceptance of users who resist changing already established procedures, thus impairing the use of some systems to their full potential. Furthermore, the need for staff training and participative management was also recognized to achieve engagement of users to digital-assisted laboratory administration.
Results of the studies that assessed the impact of implementing computerized management systems.
| System/Software | Reference | Objective | Test Groups | Method | Results |
|---|---|---|---|---|---|
| Customer Information Control System/Virtual Storage (CICS/VS) | Delorme and Cournoyer [ ] | Qualitative and quantitative evaluation of system limitations and impact on workflow and man/hour relationships | Software developers and users at the Hospital Lab (N.D.) | Qualitative evaluation of the development of integrated modules; during field testing, the workflow was accessed by the evaluation of patient entry forms, the results of sample and data processing, and the final reports. | |
| CCLMS | Selznick et al. [ ] | Test system improvement on organization and control of collections. | Cell culture specialists from 2 labs (n = 6) | Qualitative and quantitative evaluation of usability through the system usability scale (SUS) and field notes. | |
| MGEA | Anderson et al. [ ] | Assess the impact on experimental workflow for gene expression analysis. | Researchers (n = 7) | A qualitative longitudinal study. Immersion in the work environment. Interviews, observations, and field notes were coded and analyzed. | |
| LINA | Yousef et al. [ ] | Effectiveness and acceptance of an inventory system for management of oligonucleotides, strains, and cell lines | Lab staff (n = 10) | Qualitative analysis of the implementation process; quantitative evaluation of usability through the SUS. | |
| AdLIMS | Calabria et al. [ ] | Evaluate effectiveness on sample tracking in genomic studies. | Developers and potential users/clients (N.D.) | Analysis of requirements and expectations of functionalities from users/clients in terms of functionalities; qualitative analysis of the development process. | |
| LabCIRS | Dirnagl et al. [ ] | Assess the acceptability, usability of a software of risk assessment for traceability of reported cases. | Lab staff (n = 31) | Statistical and qualitative analysis of the data before and after the implementation of the tool. Online questionnaire with two questions on software usability | |
| LC Virtual Biorepository of the Antibacterial Resistance Leadership Group (ARLG) | Manca et al. [ ] | Assess the impact of the implementation of a virtual repository on the management of data and biological collections. | Customers from research labs and diagnostic companies (N.D.) | Qualitative evaluation of the efficiency of the primer bank sequences. Quantitative retrospective assessment of impacts on services provided. | |
| Quartzy | Timoteo et al. [ ] | Assess the impact of the implementation in the workflow and the perception of users at an academic laboratory. | Lab staff (n = 30) | Qualitative analysis of the team’s attitude towards implementation, including a structured questionnaire (and focus group assessments). Management performance indicators were also compared before and after implementation. | |
| SDLC | Dennert, Friedrich, and Kumar [ ] | Evaluate the development steps of a database of biological sample inventories | Researchers from different fields of medicine (N.D.) | Immersion in the work environment: The cycles of all resources have been developed and tested. User training and interviews were conducted to assess the applicability and identify user’s needs. |
N.D.: non-determined number of participants.
Regarding the management subjects issued according to each laboratory, digital systems were employed for several different uses, from purchases and administrative tasks to control of cell collections, inventories in general, as well as data storage and management of animal colonies.
All the thirty-two described software issued one or more topics of management recommended by documents of good laboratory practices [ 3 ], including experimental workflow, data storage, integration with laboratory equipment, statistical analysis, comparison of experimental data, animal colonies, biorepositories, inventory, and risks. The integration of work demands of academic health sciences laboratories and items of compliance with the GLP guidelines are identified in the chart presented in Figure 2 .

The main applications of the identified software on the different sections and chapters of the OECD GLP Principles [ 3 ].
4. Discussion
4.1. contributions to adherence to glp principles.
While the search strategy from the present review identified several different laboratory management systems, few of the eligible studies provided a focused discussion on this topic. The lack of direct scientific evidence limits the present review to quantitatively assess to what extent digital systems can collectively contribute to accreditation achievement. On the other hand, all the identified software accounted for management issues related to at least one of the GLP principles, and, in some studies, more than one software was used to meet the different demands related to quality systems.
In this sense, the approach proposed by Timoteo et al. [ 6 ] could be applied to the present sources of data to chart the main topics of management affected by these programs and systems related to good practice guidelines. The chart presented in Figure 2 shows how the types of management supported by the software in academic laboratories are related to several items from Section II of the OECD GLP Principles [ 3 ]. Such relationship is revealed by an emphasis on the responsibilities of staff and facilities management, work planning, availability of standard operational procedures (SOPs) that cover all study activities, procedure analysis, use and maintenance of equipment, as well as the application of standards for receiving test samples, its chain of custody and logistics, control of inventory, and the traceability of reagents and validation of methods.
For a better understanding of the functions of these systems, a brief presentation of them will be made, with an emphasis on meeting the computerized systems to the GLP principles listed in Figure 2 .
4.1.1. Workflow
The GLP principles require precise definitions of the different steps during the performance of the study, as described in item #8 of the OECD document [ 3 ], including the responsibilities of the personnel involved, the facilities and status of equipment employed (item #3), among other factors. Furthermore, quality assurance (item #2) requires identifying and monitoring critical steps, checkpoints, and possible sources of errors. Among the different systems identified in the present review, some described digital tools dedicated to managing such workflow of study performance in a systematized fashion.
In the late 1990s, Goodman and colleagues [ 16 ] presented Labflow, a software dedicated to genetics and mapping studies. Workflow management was not recognized as a study topic at that time and, while LIMS already existed, there was no commercial LIMS product that supported workflow management in a specific sense. In this scenario, LabFlow appeared among the first digital solutions, with a workflow model in which objects flow different laboratory tasks (such as DNA extraction, selection of clones, sequence analysis) under programmatic control. An essential point of this software was already allowing the programmer to customize their workflows to different laboratory needs.
Anderson et al. [ 19 ] described, in 2007, the implementation of the Microarray Gene Expression Analysis (MGEA), a software package developed by Rosetta Biosoftware (a subsidiary of Merck Inc.), that helped to integrate workflow information related to experimental design, data collection, and bioinformatic analysis of genomic results. Despite the high costs of the license and its renewals, the authors expected that implementing a commercially available service would bring advantages such as security in terms of support for operation and uniformity between different research centers, thus facilitating communication between employees. However, their qualitative analysis observed that the system was not used to its full potential, and its acceptance by staff would demand ongoing training and even an evolution of academic curricula towards the use of bioinformatics tools.
In 2019, Gaffney et al. [ 11 ] described the design and implementation of GEM-NET, a software that allowed members of the C-GEM (Center for Genetically Encoded Materials, USA) to integrate research efforts connecting six laboratories spread across three university campuses. GEM-NET was designed to support science and communication by integrating task management, scheduling, data sharing, and internal communications. A set of more than 20 tools was organized, including two applications customized for the Institution’s specific needs of workflow management. The tools are highly interconnected, but the set can be divided into access control, data storage, data navigation, project monitoring, teamwork, internal communication, and public engagement. The authors conclude that GEM-NET provides a high level of security and reliability in workflow management.
4.1.2. Data Management
In different items of the GLP principles, a need is described for the secure storage, filing, and retrieval of research data (item #7.4), including study plans, raw data, final reports, test system samples, and specimens (item #8.3), and their related archiving facilities (item #3.4). Furthermore, item #7 (standard operating procedures) requires the preparation and observance of documents that guarantee the quality and integrity of the data generated by the studies. Sub-item #7.4, for example, describes that in the case of computerized systems, validation, operation, maintenance, security, change control, and the backup system must be observed.
Within the selected studies, we found the report of computerized systems to manage data from various laboratory environments and how they were made available to the research groups. In the early 1980s, Delorme and Cournoyer [ 15 ], in a microbiology laboratory of a University Hospital, tested the CCIS/VS (Customer Information Control System/Virtual Storage), consisting of customer data repository, using a central computer shared with medical records databases, admission offices, patient accounting, and other medical-administrative services. The system also served as a virtual storage system, including data from microbiological samples. It performed activities such as report printing, data quality control, epidemiological assistance, germ identification, teaching, and research in the different subspecialties of microbiology. The authors carried out a qualitative and quantitative assessment identifying an improvement of workflow without increasing personnel, together with a reduction in the time for the production of reports, system downtime, and other parameters.
Viksna et al. [ 20 ] focused on collecting, storing, and retrieving data on research participants and biomedical samples through electronic management. For this, they proposed the PASSIM (Patient and Sample System for Information Management), a web-based customizable system that could be used for sending, managing, and retrieving samples and data from the research subject, ensuring the confidentiality of the records. This tool was instrumental in managing information in clinical research studies involving human beings and replaced the more expensive LIMS, which requires investments of time, effort, and resources that were not always available.
Electronic laboratory notebooks (ELN) are programs designed to replace traditional research notebooks. These electronic tools may register protocols, field/lab observations, notes, and other data inserted through a computer or mobile device, offering several advantages over paper notebooks [ 19 ]. Machina and Wild [ 22 ] investigated the importance of ELNs when integrated with other computer tools, such as laboratory information management systems, analytical instrumentation, data management systems, and scientific data. They observed that the type of laboratory (analytical, synthesis, clinical, research) was a primary source of differences when trying to integrate ELN with the available tools. Therefore, based on the observation that there was no well-established path for the effective integration of these tools, the authors decided to review and evaluate some of the adopted approaches.
Calabria et al. [ 24 ], in 2015, introduced adLIMS, a software for managing biological samples (primarily DNA) and metadata for patient samples and experimental procedures. The authors described how it was possible to produce this system by customizing a previous open-source software, ADempiere ERP. First, they collected the requirements of the end-users, verifying the desired functionalities of the system and Graphical User Interface (GUI), and then evaluated the available tools that met the desired requirements, ranging from pure LIMS to content management and corporate information systems. The authors report that the system supported critical issues of sample tracking, data standardization, and automation related to NGS (next-generation sequencing).
By 2021, Cooper et al. [ 30 ] reported using integrated systems that ensure the sharing of essential data for current research. The authors followed the 15 years of development and implementation of the LabDB system, initially projected to manage structural biology experiments, which could be improved into a sophisticated system that integrates a range of experimental biochemical, biophysical, and crystallographic data. The LabDB central software module handles data from the management of laboratory personnel, chemical stocks, and storage locations. It is currently used by the American/Canadian consortium CSGID (Center for Structural Genomics of Infectious Diseases) and several prominent research centers. The authors identified the difficulties and resistance of some researchers in adopting these systems as the main limitation, often due to the necessary effort to import data from electronic notebooks or laboratory spreadsheets, with which most researchers are already familiar. Nevertheless, the authors consider that this effort is worth it since these older approaches do not remove or even track inconsistencies and do not adapt well to the requirements of modern research.
It is essential to notice that, for accreditation purposes, hosted services (cloud archiving, backup, or processes) require written agreements describing the responsibilities of the informatics services. Test facility management must be aware of potential risks on data integrity resulting from third-party storage.
4.1.3. Equipment
Adherence to the GLP principles speaks to the adequate management of research equipment (OECD item #4), including their adequate calibration, maintenance, scheduling, and responsible staff in the test facility. Several commercially available systems, such as QRESERVE, cited by Perkel [ 9 ], are entirely dedicated to these functions, with integrated reservation calendars, administration of equipment status and availability, a repository of maintenance documentation, and a registry of use time. Other all-purpose management systems such as Labguru have most of these functions on a specific equipment module. That was also the case of the freely available (for individual researchers) Quartzy until 2016, as reported by Timóteo et al. [ 6 ]. This study described how the implementation of the software optimized the shared use of equipment on a multiuser clinical research unit and the advantages of allowing equipment scheduling, check-in, and check-out remotely, even using mobile phones.
4.1.4. Animal Facilities
Several procedures related to pre-clinical studies conducted with animals are issued in the GLP principles, mostly in item #5 of the OECD document (test system) and subsection #5.2 (Biologicals). These include a proper registry of housing, handling, and care of animal test systems to ensure the quality of the data. Additionally, records of source, date of arrival, and arrival condition of test systems should be maintained. Two selected studies described the use of vivarium monitoring software to ensure the remote control of stocking, accommodation, handling and care of animals, identification of colonies, and inventory of supplies.
Milisavljevic et al. [ 8 ] described, in 2010, the Laboratory Animal Management Assistant (LAMA), a software modified from the LIMS proposal to optimize small animal research management. It was initially developed to manage hundreds of new mouse strains generated by an extensive functional genomics program in Canada. The authors realized that they needed greater availability of suitable, easy-to-use systems and software interfaces. LAMA was implemented for a broad community of users, allowing individual research labs to track their colonies in a larger facility, independently. This open-access software is still available to the research community.
Allwood et al. [ 23 ] described, in 2015, how smartphones could help researchers in the remote management of animal colonies. The authors proposed Lennie, an app that introduced a new method for managing small to medium-sized animal colonies, allowing users to remotely access the facilities, and create and edit several functions virtually from anywhere. Its use contributes to the optimization of workflow and planning of experiments, offering a user-friendly experience. Possible updates to the functionalities were also suggested, such as camera integration with the calendar, permission for data sharing, and permanent storage.
4.1.5. Biobank/Repository
In order to comply with the GLP standards, samples that arrive at a laboratory must have records that include the characterization and reference, date of receipt, expiration date, quantities, and storage data, following item #6.1 (receiving, handling, sampling, and storage). This issue is of utmost importance for managing biobanks and biorepositories, creating a need for specific software for successful management.
Boutin et al. [ 25 ] carried out a study on a complex system of various software that contributed to the management of a Biobank. The core object of management was an extensive repository of samples and data available to researchers. The platform requires robust software and hardware, as they work with large amounts of data stored and transferred to research groups. In the study, the authors described each of the five custom and commercially available information systems integrated into the existing clinical and research systems, and discuss safety, efficiency, and challenges inherent in the construction and maintenance of this infrastructure. Constrack was used to manage patient data. The Enterprise Master Specimen Index (EMSI) is a sample indexing system, STARLIMS manages inventory, GIGPAD manages data and integrates equipment, and the Biobank Portal is the customized application that connects all the systems.
Manca et al. [ 28 ] assessed the structure of a central laboratory of the Antibacterial Resistance Leadership Group (ARLG) in the USA. This group leads the evaluation, development, and implementation of laboratory-based research and supports standard or specialized laboratory services. The laboratory included both a physical and a virtual biorepository. They developed digital procedures for reviewing and approving strain requests, providing guidance during the selection process, and monitoring the transfer of strains from the distribution laboratories to the requesting investigators.
Paul et al. [ 29 ] also describe a Biobank management system, with great emphasis on data storage in clouds. The authors evaluated that biobanks have become an essential resource for health research and drug discovery. However, collecting and managing large volumes of data (bio-specimens and associated clinical data) requires biobanks to use more advanced data management solutions. Paul and Chatterjee [ 27 ] point out that in the current COVID-19 pandemic scenario, that requires global and quick actions, virtual biobanks present a crucial role in several different fronts, from diagnosis to research. Without the need to physically use biological samples, these banks may allow sharing medical data and networks for better cooperation between biobanks at the national and international levels.
Recently, Dennert, Friedrich, and Kumar [ 1 ] explained the various implications of the inventory management of biological samples from various research areas, employing different cryopreservation methods. Such management must ensure the availability of items, easy tracking, and the optimization of shared space among the various research groups. For this, the authors presented the various stages of developing an inventory data model using the Microsoft Access database, after several phases that included training, planning, implementation, and maintenance, as well as the establishment of manuals and protocols for standardized data entry. Using the software development lifecycle (SDLC), the authors attained a database construction model. This model requires frequent communication with users to provide transparency and quality improvement.
4.1.6. Risk Management
Identifying incidents and risk assessment is an essential part of the GLP standards that requires an adequate work plan and a quality assurance program (OECD document item #2). Item #8.3 of the GLP states that all data changes during the conduction of a study must always be registered and responsible for the change to ensure traceability, enabling a complete audit trail to show all changes without masking the original data.
The work of Dirnagl et al. [ 27 ] discusses how error management is fundamental to comply with international standards while studying the implementation of the LabCIRS (Laboratory Critical Incident Reporting System), a simple, accessible, and open-source critical incident reporting system for pre-clinical and basic academic research groups. The software was implemented by establishing an electronic quality management system, which allowed accessibility through any laboratory computer, enabling incident reports that included photo uploads and automatic alerts for new reports and archiving.

4.1.7. Inventory
Item #6.2 of the GLP principles clearly states that all material from a study must be adequately identified, including the batch number, purity, composition, concentrations, or other characteristics, to define each item or reference item properly. It also indicates the need to keep the receipt and expiration dates, quantities received/used in the studies, and storage instructions for the stock of materials. In this review, several articles emphasized this need to monitor inventories with the help of computerized systems.
Nayler and Stamm [ 17 ], in 1999, described a laboratory management software, ScienceLab Database (SLD), which offered a management platform for molecular biology research laboratories. The program primarily manages the stock of biological samples, including plasmids, antibodies, cell lines, and protocols, and included an ordering and grants management system. The authors considered that this system met the specific needs of a small to medium-sized research laboratory, helping to organize inventories of valuable reagents, storing, and maintaining information about these items, and simplifying orders and processes.
By 2016, Catena et al. [ 26 ] developed the AirLab, a cloud-based tool with web and mobile interfaces, to organize antibody repositories and their multiple conjugates. Due to the large number of data generated by these collections, the authors recognized the need for dedicated software. The work demonstrated that Airlab simplifies the purchase, organization, and storage of antibodies, creating a panel to record results and share antibody validation data.
Yousef et al. [ 21 ] described the LINA (Laboratory Inventory Network Application) as a set of Windows-based inventory management software configured to work on a computer network with multiple users. Designed for small molecular biology laboratories, it uses Access databases to assign a new identifier to each new reagent, providing a library that helps with research and comparing DNA sequences. It later faced several features, such as expanding the types of tables available, compatibility with other operating systems, barcoding, and improvement of security issues. According to the authors, the resources provided by LINA are comparable to those available in commercial databases, with the advantage of providing a free database maintenance application for academic laboratories.
In an opinion article published in Nature’s section “Toolbox”, Perkel [ 9 ] describes several low-cost computerized electronic inventory systems as a means to overcome tortuous searches, old notebooks, out-of-date spreadsheets, and “frost-encrusted freezer boxes” to identify laboratory samples and resources. Besides programs discussed by other authors in this review, such as LINA and Quartzy, the article cites other systems such as OpenFreezer, a free web-based system to register sample data such as location, origin, and biological properties, the cloud-based StrainControl (DNA Globe, Sweden), a software free for individual researchers that provides support for managing different lab-organism strains, molecules, and chemicals, the mLIMS, developed by BioInfoRx (Madison, WI, USA), designed to track rodent colonies, LabGuru (BioData, Cambridge, MA, USA), a widely known paid cloud-based all-in-one Electronic Notebook, and CISPro (BioVia, Waltham, MA, USA), described as a functional Institute-wide tracking system for shared resources. Despite differences in accessibility and several resources, all of these systems share similar search engines linked to customizable databases.
Timoteo et al. [ 6 ] evaluated, by 2020, the impact of implementing a multi-module, free-of-charge online management system (Quartzy, Quartzy Inc., Santa Clara, CA, USA) in the workflow of a Brazilian academic clinical research laboratory on the perception of users. Until 2016, the software modules could assist in various aspects and demands of the laboratory, including user communications, multiuser equipment management, material inventory, research documents, and tracking of supply orders. Unfortunately, Quartzy was recently updated to a simpler version, consisting only of an inventory and purchase tracking system that connects researchers to hundreds of life sciences brands and suppliers.
4.2. Evaluating Impacts and Limitations
Effectiveness is a fundamental point to be considered in the potential role of software for laboratory management. However, most of the eligible studies identified in our search did not investigate the reported systems’ impact either through qualitative or quantitative assessments. Moreover, despite the performance of evaluations, few studies identified or discussed the limitations and drawbacks of the studied information systems. The studies with evaluations reported, among several aspects, improvement of the organization, workflow, traceability, reliability, acceptability, and good use of the software. Decreased process errors were reported that were made manually, thereby gaining productivity and reducing work. In some specific cases, they positively evaluated the control of frozen cells, generating efficiency and better results in partner laboratories. On the other hand, regarding limitations, older articles (before 2000) identified problems that were more related to system performance, which was sometimes slow and needed adjustments at a time when information technology was still incipient. The limitations from the most current systems are more related to a selective satisfaction and acceptance of software tools, specific according to the function and objective of each group and, in some cases, the resistance by researchers and staff to abandon old ways and migrate to digital tools, which were not used to their full potential within the laboratory.
To adequately assess the impact of these electronic management systems, different methodological approaches are available, such as pre/post-tests evaluating quantitative indicators of performance and provision of services. However, as Timoteo et al. [ 6 ] discussed, the complex nature of the provided services of multiuser, academic research facilities may impair the obtention of feedback through quantitative indicators. In this sense, the perception and attitudes of staff towards the management system may contribute to understanding its impact on the workflow and the search for quality at academic clinical research laboratories, as well as provide data for the development or improvement of actions and strategies toward quality and compliance [ 31 , 32 , 33 ]. In this sense, validated tools may provide a means to standardize the evaluation of laboratory management software, allowing comparisons on the effectiveness and adequacy of these systems in different applications. Two studies [ 18 , 21 ] proposed the use of an important tool to investigate the effectiveness and efficiency of the software, the system usability scale (SUS). This tool, developed by John Brooke at Redhatch Consulting (UK), consists of a simple, ten-item attitude questionnaire using a Likert scale to provide a global view of subjective assessments of usability, which was validated as providing reliable results even with small samples/study groups, which was the case of most identified studies in this review. Therefore, it may represent a potential tool (although underestimated until the present moment) for further studies on implementing laboratory management systems.
Different studies point out that staff training is one of the most important factors of success of the implementation of these systems and a key part in acceptance and adapting to a new management model. Dirnagl et al. [ 27 ] evaluated the impact on staff attitudes toward incident reporting after one year of implementation, observing that training led to greater adherence to the goal of complying with international quality standards and mature culture of error management. Timóteo et al. [ 6 ] performed a qualitative evaluation of the staff perception on software implementation, where most users stated that constant training and leadership were pivotal for the successful use of the software. On the other hand, Anderson et al. [ 19 ] reported that limited access to training was a barrier to software use during the implementation of MGEA, and that the lack of ongoing training might have contributed to a progressive de-emphasizing of the system use among the laboratory staff. These data point to the need of careful planning by the PIs to ensure continuous and inclusive training on the implementation program of management systems.
4.3. Software Availability
Regarding availability and accessibility, until 2010, most of the identified programs had to be downloaded/installed to specific laboratory computers [ 19 , 30 ], but were sometimes able to integrate local area networks (LANs), as described by Delorme and Cournoyer in 1980 [ 15 ]. In the past decade, technology has advanced to online software, expanding even to applications (apps) on mobile phones, reflecting the current expectations of users and consumers. With app technology permeating all fields of our daily lives, it would be natural for this technological paradigm to reach laboratory and research technologies. Indeed, a big leap was identified towards the proper integration between lab management systems and the new mobile universe. Real-time communication makes it possible, for example, that inventory checks, equipment scheduling, and data verification of an animal colony be performed while in transit. Multicenter studies can share data in real-time, as recently observed in the fast development studies of vaccines against SARS-CoV-2 since 2020, relying heavily on technological development and efficient data management [ 34 ].
Begg et al. [ 35 ] discussed how computer systems are of particular importance in the process of GLP certification in low- and middle-income countries, even though their role is not always emphasized on accreditation systems around the world. This review identified that the knowledge on laboratory management software is mainly originated, as expected, from developed, high-income countries, with advanced information technology industries and significant investment in technology and support for universities and study centers (USA, Germany, Canada, United Kingdom, Switzerland). In a critical view, it may indicate an economic bias in the technological development on the theme, as developing countries maintain a role as consumers of technology and not as producers and developers, reflecting little investment in this (and other) technological areas.
The costs of implementing computerized systems may represent one of the main challenges for public Academic Health Centers since these Institutions, in general, face tight budgets to support several laboratories, researchers, and research lines. Such limitations are expected to be potentialized when considering low- to middle-income countries, which could benefit from low-cost or cost-free initiatives.
In general, the development and maintenance of information systems are made possible by providing subscription services to ensure the tool’s sustainability. The present review identified some systems that addressed a full spectrum of fundamental issues in the management of academic laboratories, such as inventory control and organization and equipment scheduling, on a free-of-charge basis, as it incorporated catalogs from various sponsors (reagent suppliers) and suggests these products when orders are placed [ 9 ]. However, such a business model probably did not match the maintenance costs of the platform, as Quartzy has shut down all functions not related to inventory/purchases by 2016, and recently included a fee for Institutional users. It is also possible that users from outside the USA and Europe could not use the vendor-related functionalities, as customer services and representatives in regions such as South America would not connect directly to the system [ 6 ]. On the other hand, LINA is an example of a system that could remain free-of-charge, even though limited to the needs of small molecular biology laboratories [ 21 ], with much simpler functionalities compared to well-known commercial applications such as Labguru. Other services, such as QReserve, have both free and paid versions with increased functionalities, allowing low-budget academic laboratories to use some free resources, such as equipment reservation and management, through a more straightforward interface.
A usual profile among entirely free software originates from in-house academic software, such as Biobank Portal and CCLMS, customized for the personal use of the developer group, usually without widespread use in other institutions. Even though they may present advantages on issuing specific demands of developers, the lack of a profound, systematic evaluation of performance on most selected studies does not allow to infer whether these are more or less effective than commercial software. In this sense, Boutin et al. [ 25 ] report that the laboratory IT framework may face challenges common to industry settings, where cost-overrun is prevented by planning the cost-effectiveness of purchasing commercially available vs. designing in-house custom applications. An interesting way to achieve broader applicability for such software is to use open-source codes, such as Boutin et al. [ 25 ], paving the way for other programmers to adapt the tool to different laboratory specificities. It is important to notice that investments from government bodies worldwide could also contribute to the development of freely available tools as part of public policies focused on increasing overall quality and adherence to good practices in health sciences research. In this sense, the encouragement of startups involving interdisciplinary initiatives can turn universities and academic centers into important stakeholders in covering technological gaps in low- or middle-income countries [ 36 ].
4.4. Review Limitations
The present Scoping Review has limitations mainly related to the impossibility of exhausting the literature on laboratory software, reflected in the choice of not including programs that dealt only with the transmission and handling of analysis results and laboratory data, such as pure LIMS or analytical bioinformatics software. Despite their fundamental role, these types of software have already been widely discussed [ 37 , 38 , 39 , 40 ], and most of these systems were not designed to support the management of staff and shared resources, for example. Additionally, the scientific literature probably does not reflect the abundance of available software since developers and the scientific community usually treat them as a commercial tool rather than a research topic. Nevertheless, regardless of such limitations, the present review was able to map a framework that points to the great applicability of these systems in the search for quality and good practices in academic experimental medicine laboratories, where restrictions regarding the availability of resources and staff and limited management experience are common restrictions. Therefore, the gaps identified here can serve as an indication for new studies that seek to assess, quantitatively or qualitatively, the impact of implementing these tools on the best practices at academic health Institutions.
5. Conclusions
The present literature review mapped several studies in the last four decades, proposing and evaluating the impact of digital tools in the management of health sciences research laboratories to several different applications, ranging from administrative workflow management and data traceability to virtual biobanking. These functions have the potential to contribute to the adherence to different GLP principles. However, the evidence for their effectiveness is still limited and requires further investigative efforts.
Acknowledgments
The authors acknowledge the financial support in scholarships from the Brazilian agencies CNPq, CAPES, and FAPERJ. The authors acknowledge the technical support by Jean Carlos Nascimento.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/healthcare9060739/s1 , Table S1: Preferred Reporting Items for Systematic reviews and Meta-Analyses extension for Scoping Reviews (PRISMA-ScR) Checklist.
Author Contributions
Conceptualization, G.A., M.T. and B.O. (Bruna Oliveira); methodology, G.A. and C.F.d.A.B.M.; software, P.M.; formal analysis, M.T., R.B., J.d.S. and L.D.; investigation, M.T., E.L., J.d.S. and G.A.; resources, P.M. and C.F.d.A.B.M.; data curation, C.F.d.A.B.M. and G.A.; writing—original draft preparation, M.T., E.L., A.C.B. and L.D.; writing—review and editing, G.A. and C.F.d.A.B.M.; project administration, B.O. (Beni Olej). All authors have read and agreed to the published version of the manuscript.
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Data availability statement, conflicts of interest.
The authors declare no conflict of interest.
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Computer Science and Engineering
- Current Students
- Faculty Directory
- Faculty Research
- PhD Admissions
Research in Computer Science and Engineering
New tool helps predict progression of alzheimer’s.
UTA’s framework can pinpoint clinical status of individuals within the disease spectrum
Augmented reality tech would help maintain military aircraft
UTA researcher earns grant to integrate computer vision into Air Force maintenance training
Supercomputing training at Argonne National Laboratory
UTA postdoc attends intensive program that includes hands-on sessions with supercomputers
Secure networks for battlefield communications
UTA researcher working to help military use civilian 5G networks securely
‘I’m not going to take it for granted’
UTA doctoral student earns prestigious scholarship from Department of Defense

Research Labs in Computer Science and Engineering
Our department has a number of teaching and research laboratories, centers and other facilities that support its educational and teaching mission. Its internationally recognized faculty members are engaged in breakthrough research across the leading areas of computer science and engineering.
Abacus Cloud and Edge Systems Lab (ACES)
Mission: At the ACES Lab, we focus on modeling, analysis, and resource management for large-scale parallel and distributed computing systems. In particular, we are interested in developing highly scalable, user-centric resource allocation solutions for cloud and edge computing. To this end, user-utility-based, distributed, joint compute-network-and-storage resource allocation solutions with provable convergence and optimality properties are sought. [Lead Faculty: Hong Jiang, Hao Che]
Adaptive and Scalable Systems Lab
Mission: The Adaptive and Scalable Systems Lab focuses on building computer systems that are adaptive to changing workloads, scalable for platform growth, and capable of providing quality-of-service guarantees and service differentiation. We combine performance analysis at the application, OS, and hardware levels with machine learning techniques to characterize the complex behaviors of computer systems. [Lead Faculty: Jia Rao]
Arlington Computational Linguistics Lab (ACL2)
Mission: This lab seeks to advance knowledge on natural language processing and knowledge engineering, with particular interest in the interdisciplinary research that empowers other scientific fields. [Lead Faculty: Kenny Zhu]
ASSIST Laboratory
Mission: The ASSIST Laboratory's focus is on researching and developing technologies to assist the elderly and people with disabilities in their everyday life. [Lead Faculty: Farhad Kamangar, Manfred Huber, David Levine, Jason Losh]
Autonomous and Intelligent Systems Lab
Mission: The Autonomous and Intelligent Systems Lab operates in partnership with the Automation and Intelligent Systems Division of the UT Arlington Research Institute’s Division of Automation and Intelligent Systems. We conduct research in machine learning, robotics and autonomous systems to solve real-world problems. Focuses include navigation and control algorithms for a wide variety of autonomous vehicles, assistive robotics for people with disabilities and mobility impairments, computer vision for environmental monitoring, and learning from demonstration for safe cooperative robotics. [Lead Faculty: Nicholas Gans]
Biomedical Computing and intelligent Systems Lab (BioMeCIS)
Mission: Developing efficient algorithms to solve computational problems in basic medicine and clinics, while making theoretical and fundamental contributions to machine learning, data mining, pattern recognition, and computer vision. [Lead Faculty: Jean Gao]
Mission: At Cyber Guard Research Lab, our mission is to safeguard emerging platforms, medical applications, web frameworks and other widely-used technologies by uncovering software vulnerabilities and developing robust defense solutions. Through the utilization of cutting-edge tools powered by program analysis, machine learning, natural language processing, and LLM, we diligently identify potential threats and propose innovative strategies to fortify the cyber-physical ecosystem. Our commitment lies in ensuring the safety and security of these platforms, ultimately contributing to a more resilient digital landscape. [Lead Faculty: Faysal Hossain Shezan]
Data Security and Privacy Lab
Mission: Develop novel methods to enable information security and privacy in modern communication systems that are robust against computationally unbounded adversaries, with a focus on solutions that are resistant against quantum-capable adversaries. [Lead Faculty: Remi Chou]
Database Exploration Lab (DBXLab)
Mission: At Database Exploration Lab (DBXLab), we seek to investigate fundamental research issues arising in Big Data. Our research encompasses diverse areas such as data mining, information retrieval, data uncertainty and probabilistic methods, approximate query processing, data summarization, data analytics and data exploration of hidden web databases, social and collaborative media. [Lead Faculty: Gautam Das]
Digital Design Laboratory
Mission: The DDL supports the design and prototyping of digital systems for educational and special purpose applications employing legacy and current design tools and implementation technologies. [Lead Faculty: Bill Carroll]
Health Data Science Lab (HDSL)
Mission: building computational tools and frameworks that at massive scale allow for cancer imaging data to be 1) contextualized in the oncology clinic to improve patient outcomes and 2) leveraged at the bench to augment drug discovery efforts. The lab also focuses on developing computational and statistical methods for handling high throughput `omics data such as single cell transcriptomics/spatial transcriptomics (10X Visium & Chromium), spatial proteomics (CODEX), and calcium imaging. [Lead Faculty: Jacob Luber]
Human Centered Computing
Mission: The mission of the Heracleia (heracleia.uta.edu) lab computational innovations in the areas of Human Computer Interaction (HCI), Pervasive Computing, healthcare services for disabilities, data analytics for behavior monitoring applications, assistive robotics, and computer aided rehabilitation. [Lead Faculty: Fillia Makedon]
Hybrid Atelier
Mission: The Hybrid Atelier is a creative technology research makerspace, serving as a nexus between the Arts, Engineering, and Sciences, with the mission of re-imagining, inventing, and supporting what creativity and making will look like 20 years from now. The atelier focuses its efforts in the areas of Human-Computer Interaction, Design, Physical Computing, Digital Fabrication, and Augmented Environments. [Lead Faculty: Cesar Torres]
Immersive Efficient Computing and Communication Lab
Mission: The Immersive Efficient Computing and Communication (IMEC 2 ) lab is dedicated to developing efficient technologies for future immersive computing and communication. Specifically, we have been focusing on mobile systems and networks, including virtual, augmented, and mixed reality (VR, AR, and MR), digital twins, etc., over 4G/5G and beyond. Our overarching mission is to enable technologies for mobile systems and networks in another decade, extending beyond form factors such as VR/AR headsets. [Lead Faculty: Jiayi Meng]
Information Technology Laboratory (IT Lab)
Mission: The mission of this lab can be summarized as: a) carry out both fundamental and practically applicable research & development, b) interact and collaborate with industry/federal agencies for identifying fundamental problems, and c) provide a viable migration path for integrating new techniques/solutions into real-world systems/applications. [Lead Faculty: Sharma Chakravarthy]
Innovative Data Intelligence Research (IDIR) Laboratory
Mission: The Innovative Data Intelligence Research (IDIR) Laboratory conducts research in several areas related to big data intelligence and data science, including data management, data mining, natural language processing, applied machine learning, and their applications in computational journalism. The lab's current research focuses on building large-scale human-assisting and human-assisted data and information systems with high usability, high efficiency and applications for social good. Particularly, the ongoing research projects include data-driven fact-checking, exceptional fact finding, fake-news detection, usability challenges in querying and exploring graph data, knowledge databases, and data exploration by ranking (top-k), skyline and preference queries. The lab started the inter-disciplinary research in computational journalism at UTA in 2010 and has since been at the frontier of this nascent field. [Lead Faculty: Chengkai Li]
Machine Learning and Computer Vision for Clinical Applications
Mission: We focus on customizing machine learning and computer vision techniques to solve various clinical problems including computer aided diagnosis using multi-modal and multi-source datasets such as medical imaging, EHR and clinical reports data collected from different institutes or hospitals; smart patients surveillance system using multiple censoring dataset collected from cameras, wifi and radar. [Lead Faculty: Yingying Zhu]
Medical Imaging and Neuroscientific Discovery Laboratory
Mission: Our research in MIND mainly focuses on the discovery of fundamental principles of brain structural and functional architectures and their relationship, via brain imaging, computational modeling and machine learning methods. We are interested in the interaction between Artificial Intelligence (AI) and Human Intelligence (HI): Using Deep Learning to facilitate the analysis and interpretation of brain data; Applying neuroscience knowledge to design more efficient Deep Learning architectures. We also have strong interests in applying the discovered principles, theories and methods to better understand neurodevelopmental, neurodegenerative and psychiatric disorders including Autism, Alzheimer’s disease, and Major Depression, among other brain conditions. [Lead Faculty: Dajiang Zhu]
Mobile Computing and Security Lab (MobiSec)
Mission: The mission of the lab is to develop effective mechanisms to address security/privacy challenges, improve system performances, and explore novel applications in mobile computing. [Lead Faculty: Ming Li]
Mobile Machine Intelligence Lab (Noodle Lab)
Mission: Our primary objective is to conduct rigorous research and development focusing on integrating software and hardware co-design principles in the AI ecosystem. The aim is to create trustworthy, efficient, and adaptable AI models that can operate effectively within the limited resources of on-device platforms, then democratizing AI to real-world on-device applications. Our research draws upon methodologies from mathematical tools, machine learning, computer architecture, and high-performance computing. We specifically build the general framework of software/hardware co-design for efficient and trustworthy AI based on higher-order tensor decomposition and optimization algorithms, then apply the corresponding AI models to on-device applications. [Lead Faculty: Miao Yin]
Rigorous Design Lab (RiDL)
Mission: Today’s computer systems have been continuously evolving to catch up with the demands of modern society. The technological progress is stretching the boundaries of what is possible, creating new unprecedented operational challenges. To that end, we focus on enhancing computer systems with secure and efficient designs through rigorous investigation and evaluation. [Lead Faculty: Mohammad Atiqul Islam]
Robotic Vision Laboratory (RVL)
Mission: The focus of the RVL is on the challenge of applying computer vision to robotics and automation. We believe that the ability to visually perceive, understand, and respond to the complex world around us is crucial for the next generation of robots in manufacturing, transportation, construction, infrastructure inspection, environmental monitoring, agriculture, healthcare, space exploration, defense, and the home. [Lead Faculty: William Beksi]
Scalable Modeling & Imaging & Learning Lab (SMILE)
Mission: In SMILE, we focus on developing scalable models and algorithms for data-intensive applications in high performance computing. In particular, we focus on advanced algorithms, software and systems for statistical learning, imaging informatics and computer vision. Our interest is to develop efficient algorithms with nice theoretical guarantees to solve practical problems involved large scale data. [Lead Faculty: Junzhou Huang]
Security and Privacy Research Lab
Mission: In this lab, we conduct data-driven research to understand online security, privacy and safety, and to develop novel techniques and frameworks for improving them. Our research is interdisciplinary and spans widely across multiple areas, from data science and social computing research to traditional security and privacy research. [Lead Faculty: Shirin Nilizadeh]
Software Engineering Research Center (SERC)
Mission: The mission of the Software Engineering Research Center (SERC) is to conduct cutting-edge research in various areas of software engineering, including software design, specification, analysis, verification, and testing. [Lead Faculty: Christoph Csallner, David C. Kung, Yu Lei, Allison Sullivan]
Transformative Wireless Systems and Technology (TWiST) Lab
Mission: At the Transformative Wireless Systems and Technology (TWiST) Lab, our commitment is to lead the way in pioneering groundbreaking advancements in wireless communication and systems. Through innovative research initiatives spanning areas such as Networked Robotics, Spectrum Learning, Open-RAN, and beyond, we aspire to redefine the capabilities of wireless systems. Our multidisciplinary team operates at the forefront of technology, integrating AI, ML, AR/VR, and other cutting-edge fields to develop transformative solutions. With a steadfast focus on practical applications and tangible real-world impact, we endeavor to shape the future of wireless technology and facilitate its seamless integration with intelligent robotics. [Lead Faculty: Debashri Roy]
Vision-Learning-Mining Research Lab (VLM)
Mission: The VLM lab is a research lab at the Computer Science and Engineering Department of the University of Texas at Arlington. At the VLM lab we are conducting research in the areas of computer vision, machine learning, and data mining. Areas of focus include gesture and sign language recognition, human motion analysis, detection and tracking of complex shapes, large-scale multiclass recognition, and similarity-based retrieval and classification using large databases. [Lead Faculty: Vassilis Athitsos]
Wireless Networks and Systems Lab (WINS)
Mission: The WINS lab, directed by Dr. Yonghe Liu, focuses on challenging issues in wireless networks. The group's research spans both theoretical study and practical system design and development. Our current research projects include novel architecture for sensor networks, routing and buffer management for delay tolerant networks, networking issues in opportunistic networks, and mobile social networks. [Lead Faculty: Yonghe Liu]
Research Areas
Our faculty conduct research in six general areas: Artificial Intelligence, Big Data and Data Science, Computer and Network Systems, Human-Computer Interfaces, Security, and Software Engineering. Each general area is divided into more specific focus areas.
Undergraduate Applicants
- Let Us Know About You
- Undergraduate Catalog
- Academic Advisors
Graduate Applicants
- M.S. Admission Requirements
- Ph.D. Admission Requirements
- Request Information
- Grad Advisors
- GAANN Fellowships
Graduate Certificates
- Certificates in Catalog
Mail & Deliveries
Box 19015 Arlington, TX 76019
CSE Department c/o UTA Central Receiving Box 19015 1225 West Mitchell Arlington, TX 76019
Phone, Fax, & Apps
- Phone: 817-272-3785
- Fax: 817-272-3784
- Student Apps
- Faculty Apps
- Research & Faculty
- Offices & Services
- Information for:
- Faculty & Staff
- News & Events
- Contact & Visit
- About the Department
- Message from the Chair
- Computer Science Major (BS/BA)
- Computer Science Minor
- Data Science and Engineering Minor
- Combined BS (or BA)/MS Degree Program
- Intro Courses
- Special Programs & Opportunities
- Student Groups & Organizations
- Undergraduate Programs
- Undergraduate Research
- Senior Thesis
- Peer Mentors
- Curriculum & Requirements
- MS in Computer Science
- PhD in Computer Science
- Admissions FAQ
- Financial Aid
- Graduate Programs
- Courses Collapse Courses Submenu
- Research Overview
- Research Areas
- Systems and Networking
- Security and Privacy
- Programming Languages
- Artificial Intelligence
- Human-Computer Interaction
- Vision and Graphics
- Groups & Labs
- Affiliated Centers & Institutes
- Industry Partnerships
- Adobe Research Partnership
- Center for Advancing Safety of Machine Intelligence
- Submit a Tech Report
- Tech Reports
- Tenure-Track Faculty
- Faculty of Instruction
- Affiliated Faculty
- Adjunct Faculty
- Postdoctoral Fellows
- PhD Students
- Outgoing PhDs and Postdocs
- Visiting Scholars
- News Archive
- Weekly Bulletin
- Monthly Student Newsletter
- All Public Events
- Seminars, Workshops, & Talks
- Distinguished Lecture Series
- CS Colloquium Series
- CS + X Events
- Tech Talk Series
- Honors & Awards
- External Faculty Awards
- University Awards
- Department Awards
- Student Resources
- Undergraduate Student Resources
- MS Student Resources
- PhD Student Resources
- Student Organization Resources
- Faculty Resources
- Postdoc Resources
- Staff Resources
- Purchasing, Procurement and Vendor Payment
- Expense Reimbursements
- Department Operations and Facilities
- Initiatives
- Student Groups
- CS Faculty Diversity Committee
- Broadening Participation in Computing (BPC) Plan
- Northwestern Engineering

Research Groups & Labs
Jump to a Section
AquaLab - Research in Distributed Computing
Artificial intelligence group, assistive and rehabilitation robotics laboratory, campanoni lab, cognition, creativity and communication (c3), habits lab - mobile health and passive sensor analytics, ideas - design automation of intelligent systems, interactive audio lab.
- Northwestern Lab for Internet and Security Technology
Midwest Uncertainty Collective Lab
Northwestern networks group.
- Northwestern Security & Privacy Group
- NuLogiCS group
Parallel Architecture Group at Northwestern
Prescience lab, qualitative reasoning group, swarm robotics lab, tangible interaction design and learning (tidal) lab, technological innovations for inclusive learning & teaching (tiilt), northwestern cs theory group, vlsi research lab.
The AquaLab conducts research on large-scale distributed systems and networking. Its approach is mainly experimental, focusing on the design, deployment, and evaluation of systems that people use. This lab is headed by Fabián Bustamante .
Return to Top
Artificial Intelligence (AI) research explores the nature of intelligence and the ways in which computation can be used to explain and engineer it. Work in AI combines the scale afforded by machine learning with the expressive and organizational power of semantic information-processing and knowledge-based reasoning. Our faculty research programs include work in machine learning, cognitive modeling, language understanding and generation, planning and reasoning, robotics and human-robot interaction, computational journalism, social media analysis, computer audition, computers and education, computational creativity, and legal reasoning.
The Assistive and Rehabilitation Robotics Laboratory strives to advance human ability through robotics autonomy by easing the burden of controlling assistive machines. This lab is headed by Brenna Argall .
Research focuses on code compilation challenges for both energy efficiency and performance on commodity processors. These challenges are addressed by co-designing compilers, the computer architecture of the platform they target and programming languages. This lab is headed by Simone Campanoni .
The C3 Lab is focused on bridging the gap between the wealth of data that surrounds us and the information that is hidden within it. Using the tools of natural language processing, data analytics, and cognitive modeling the C3 Lab builds systems that link people to the information and insight that serves them. C3 is led by Kristian Hammond .
The Delta Lab is an interdisciplinary research lab and design studio at Northwestern University. Its driving mission is to improve the way we design, work, learn, play, and fundamentally change the way we interact. This lab is headed by Haoqi Zhang.
We design, build and analyze end-to-end mobile health systems, while focusing on signal processing and machine learning of wearable sensor data to help answer health-related questions. This lab is headed by Nabil Alshurafa.
IDEAS Lab aims to develop innovative design automation methodologies and algorithms for software synthesis of cyber-physical systems (CPS), which have applications in key sectors such as automotive, aerospace, healthcare, and industrial automation.
The Interactive Audio Lab applies machine learning, signal processing, natural language processing, and database search techniques to make new auditory tools and interfaces. This lab is headed by Bryan Pardo .
We are a research lab working at the intersection of information visualization and uncertainty communication. Our mission is to combat misinterpretations and overconfidence in data by developing visual representations and human-in-the-loop tools that express uncertainty and align with how people think. Topics we like include sampling-oriented uncertainty visualizations, eliciting and modeling beliefs about data, automated visualization reasoning and generation, and Bayesian statistics. The MU Collective is directed by Jessica Hullman and Matt Kay .
Return to Top
Northwestern Lab for Internet and Security Technology
The Northwestern Lab for Internet and Security Technology works to improve the reliability, agility, and security of the current Internet, including wireless networks. This lab is headed by Yan Chen .
The Northwestern Networks Group conducts research on computer networking. This lab is headed by Aleksandar Kuzmanovic .
Northwestern Security and Privacy Group
Northwestern Security and Privacy Group has a group of faculty & researchers working on software/system security, network security, AI security, cryptography, and data privacy.
NuLogiCS Group
Human intelligence lies on the capability of thinking. With the development of the programmable computer, slow thinking has found its solid foundation on computation. With the recent progress of machine learning, especially deep learning, fast thinking is discovering its root on learning. Logics is the foundation of both computation and learning. NuLogiCS group's goal is to build the new logic foundation for Artificial Intelligence (which needs a tight integration between computation and learning), and for Security (which needs a tight integration between cryptography and formal verification).
The Parallel Architecture Group at Northwestern (PARAG@N, pronounced “ paragon ”) conducts research in energy-efficient high-performance parallel computing at all scales through cross-layer design: from emerging devices and circuits, to computer architecture, compilers, runtimes, operating systems, and applications. This lab is headed by Nikos Hardavellas.
The Prescience Lab conducts a range of experimental computer systems research with a current focus on virtualization, empathic systems, and parallel and distributed systems. This lab is headed by Peter Dinda .
The research of this group includes efforts to both create new kinds of cognitive systems and model human cognition. It has strong collaborations with other cognitive scientists in a number of fields at various institutions. This group is headed by Kenneth D. Forbus .
Research interests interests include advancing the control and design of multi-robot systems, enabling their use instead of traditional single robots and to solve problems for which traditional robots are not suitable. Using these multi-robot systems can offer more parallelism, adaptability, and fault tolerance when compared to a traditional single robot. The lab is interested in investigating how new technologies will allow for more capable multi-robot systems and how these technologies impact the design of multi-robot algorithms, especially as these systems begin to number in the hundreds, thousands, or even millions of robots. This lab is headed by Michael Rubenstein .
TIDAL Lab is a team of designers, artists, learning scientists, and computer scientists at Northwestern University. Its research creates and studies innovative technology-based learning experiences. This lab is headed by Michael Horn .
The Technological Innovations for Inclusive Learning and Teaching (TIILT) Lab aims to improve learning opportunities for students from under-served communities. This lab is headed by Marcelo Worsley .
Theoretical computer science looks at fundamental questions about computation by creating formal models of computation and understanding the resources needed to solve general and specific algorithmic questions. TCS studies the design of efficient algorithms and understanding the computational complexity of various computational tasks that arise in computer science, statistics, economics and the other sciences.
The major research areas include design and analysis of algorithms, computational complexity, randomness in computation, combinatorial optimization, approximation algorithms, online algorithms. The theory group at Northwestern also has strong interests in using computation as a fundamentally new lens to study other fundamental sciences, leading to areas of algorithmic game theory, machine learning and bioinformatics.
The VLSI Research Lab at NU focuses on the areas of energy efficient computing with digital and mixed signal design approaches. We are dedicating our effort to explore new computing method for many emerging applications including energy efficient edge computing, accelerator designs for artificial intelligence, etc.
More in this section
- Engineering Home
- CS Department
Related Links
- Research at McCormick
- Meet our Faculty
- Northwestern Research Overview
Contact Info
Larry Birnbaum Professor Phone: 847-481-5258 Email Larry

The Computer Systems Research Laboratory is a research group made up of faculty members and students in the Department of Computer Science and Engineering at the University of North Texas. While we are interested in many topics within computer science, our current research focuses mainly on 3D RAM design, Processing-in-Memory and memory analysis tools. We are also investigating hardware and system level security enhancements, ontologies for capturing and analyzing security vulnerabilities and hardware assistance for managing complex indexing for neural networks, graph analytics and sparse data.
Suggestions or feedback?
MIT News | Massachusetts Institute of Technology
- Machine learning
- Social justice
- Black holes
- Classes and programs
Departments
- Aeronautics and Astronautics
- Brain and Cognitive Sciences
- Architecture
- Political Science
- Mechanical Engineering
Centers, Labs, & Programs
- Abdul Latif Jameel Poverty Action Lab (J-PAL)
- Picower Institute for Learning and Memory
- Lincoln Laboratory
- School of Architecture + Planning
- School of Engineering
- School of Humanities, Arts, and Social Sciences
- Sloan School of Management
- School of Science
- MIT Schwarzman College of Computing
Modular, scalable hardware architecture for a quantum computer
Press contact :, media download.

*Terms of Use:
Images for download on the MIT News office website are made available to non-commercial entities, press and the general public under a Creative Commons Attribution Non-Commercial No Derivatives license . You may not alter the images provided, other than to crop them to size. A credit line must be used when reproducing images; if one is not provided below, credit the images to "MIT."

Previous image Next image
Quantum computers hold the promise of being able to quickly solve extremely complex problems that might take the world’s most powerful supercomputer decades to crack.
But achieving that performance involves building a system with millions of interconnected building blocks called qubits. Making and controlling so many qubits in a hardware architecture is an enormous challenge that scientists around the world are striving to meet.
Toward this goal, researchers at MIT and MITRE have demonstrated a scalable, modular hardware platform that integrates thousands of interconnected qubits onto a customized integrated circuit. This “quantum-system-on-chip” (QSoC) architecture enables the researchers to precisely tune and control a dense array of qubits. Multiple chips could be connected using optical networking to create a large-scale quantum communication network.
By tuning qubits across 11 frequency channels, this QSoC architecture allows for a new proposed protocol of “entanglement multiplexing” for large-scale quantum computing.
The team spent years perfecting an intricate process for manufacturing two-dimensional arrays of atom-sized qubit microchiplets and transferring thousands of them onto a carefully prepared complementary metal-oxide semiconductor (CMOS) chip. This transfer can be performed in a single step.
“We will need a large number of qubits, and great control over them, to really leverage the power of a quantum system and make it useful. We are proposing a brand new architecture and a fabrication technology that can support the scalability requirements of a hardware system for a quantum computer,” says Linsen Li, an electrical engineering and computer science (EECS) graduate student and lead author of a paper on this architecture.
Li’s co-authors include Ruonan Han, an associate professor in EECS, leader of the Terahertz Integrated Electronics Group, and member of the Research Laboratory of Electronics (RLE); senior author Dirk Englund, professor of EECS, principal investigator of the Quantum Photonics and Artificial Intelligence Group and of RLE; as well as others at MIT, Cornell University, the Delft Institute of Technology, the U.S. Army Research Laboratory, and the MITRE Corporation. The paper appears today in Nature .
Diamond microchiplets
While there are many types of qubits, the researchers chose to use diamond color centers because of their scalability advantages. They previously used such qubits to produce integrated quantum chips with photonic circuitry.
Qubits made from diamond color centers are “artificial atoms” that carry quantum information. Because diamond color centers are solid-state systems, the qubit manufacturing is compatible with modern semiconductor fabrication processes. They are also compact and have relatively long coherence times, which refers to the amount of time a qubit’s state remains stable, due to the clean environment provided by the diamond material.
In addition, diamond color centers have photonic interfaces which allows them to be remotely entangled, or connected, with other qubits that aren’t adjacent to them.
“The conventional assumption in the field is that the inhomogeneity of the diamond color center is a drawback compared to identical quantum memory like ions and neutral atoms. However, we turn this challenge into an advantage by embracing the diversity of the artificial atoms: Each atom has its own spectral frequency. This allows us to communicate with individual atoms by voltage tuning them into resonance with a laser, much like tuning the dial on a tiny radio,” says Englund.
This is especially difficult because the researchers must achieve this at a large scale to compensate for the qubit inhomogeneity in a large system.
To communicate across qubits, they need to have multiple such “quantum radios” dialed into the same channel. Achieving this condition becomes near-certain when scaling to thousands of qubits. To this end, the researchers surmounted that challenge by integrating a large array of diamond color center qubits onto a CMOS chip which provides the control dials. The chip can be incorporated with built-in digital logic that rapidly and automatically reconfigures the voltages, enabling the qubits to reach full connectivity.
“This compensates for the in-homogenous nature of the system. With the CMOS platform, we can quickly and dynamically tune all the qubit frequencies,” Li explains.
Lock-and-release fabrication
To build this QSoC, the researchers developed a fabrication process to transfer diamond color center “microchiplets” onto a CMOS backplane at a large scale.
They started by fabricating an array of diamond color center microchiplets from a solid block of diamond. They also designed and fabricated nanoscale optical antennas that enable more efficient collection of the photons emitted by these color center qubits in free space.
Then, they designed and mapped out the chip from the semiconductor foundry. Working in the MIT.nano cleanroom, they post-processed a CMOS chip to add microscale sockets that match up with the diamond microchiplet array.
They built an in-house transfer setup in the lab and applied a lock-and-release process to integrate the two layers by locking the diamond microchiplets into the sockets on the CMOS chip. Since the diamond microchiplets are weakly bonded to the diamond surface, when they release the bulk diamond horizontally, the microchiplets stay in the sockets.
“Because we can control the fabrication of both the diamond and the CMOS chip, we can make a complementary pattern. In this way, we can transfer thousands of diamond chiplets into their corresponding sockets all at the same time,” Li says.
The researchers demonstrated a 500-micron by 500-micron area transfer for an array with 1,024 diamond nanoantennas, but they could use larger diamond arrays and a larger CMOS chip to further scale up the system. In fact, they found that with more qubits, tuning the frequencies actually requires less voltage for this architecture.
“In this case, if you have more qubits, our architecture will work even better,” Li says.
The team tested many nanostructures before they determined the ideal microchiplet array for the lock-and-release process. However, making quantum microchiplets is no easy task, and the process took years to perfect.
“We have iterated and developed the recipe to fabricate these diamond nanostructures in MIT cleanroom, but it is a very complicated process. It took 19 steps of nanofabrication to get the diamond quantum microchiplets, and the steps were not straightforward,” he adds.
Alongside their QSoC, the researchers developed an approach to characterize the system and measure its performance on a large scale. To do this, they built a custom cryo-optical metrology setup.
Using this technique, they demonstrated an entire chip with over 4,000 qubits that could be tuned to the same frequency while maintaining their spin and optical properties. They also built a digital twin simulation that connects the experiment with digitized modeling, which helps them understand the root causes of the observed phenomenon and determine how to efficiently implement the architecture.
In the future, the researchers could boost the performance of their system by refining the materials they used to make qubits or developing more precise control processes. They could also apply this architecture to other solid-state quantum systems.
This work was supported by the MITRE Corporation Quantum Moonshot Program, the U.S. National Science Foundation, the U.S. Army Research Office, the Center for Quantum Networks, and the European Union’s Horizon 2020 Research and Innovation Program.
Share this news article on:
Related links.
- Quantum Photonics and AI Laboratory
- Terahertz Integrated Electronics Group
- Research Laboratory of Electronics
- Microsystems Technology Laboratories
- Department of Electrical Engineering and Computer Science
Related Topics
- Computer science and technology
- Quantum computing
- Electronics
- Semiconductors
- Electrical Engineering & Computer Science (eecs)
- National Science Foundation (NSF)
Related Articles

Scaling up the quantum chip
Quantum sensing on a chip

Toward mass-producible quantum computers
Previous item Next item
More MIT News

Symposium highlights scale of mental health crisis and novel methods of diagnosis and treatment
Read full story →

Just thinking about a location activates mental maps in the brain

Nancy Kanwisher, Robert Langer, and Sara Seager named Kavli Prize Laureates

Researchers use large language models to help robots navigate
Making climate models relevant for local decision-makers
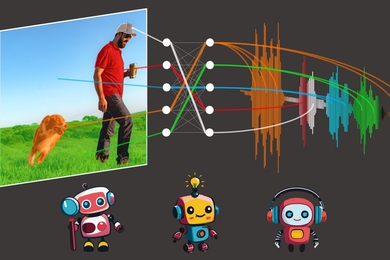

New algorithm discovers language just by watching videos
- More news on MIT News homepage →
Massachusetts Institute of Technology 77 Massachusetts Avenue, Cambridge, MA, USA
- Map (opens in new window)
- Events (opens in new window)
- People (opens in new window)
- Careers (opens in new window)
- Accessibility
- Social Media Hub
- MIT on Facebook
- MIT on YouTube
- MIT on Instagram

Did You Know?

Featured Events
Caltech's 130th commencement ceremony, astronomy on tap, behind the book: olympian and caltech athletics director betsy mitchell discusses her book.
VIEW ALL UPCOMING EVENTS
Life at Caltech

Jet Propulsion Laboratory
Nasa launches second small climate satellite to study earth's poles link opens in a new tab.
Data from the pair of CubeSats will offer new insights into how much heat the Arctic and Antarctica radiate into space.

- Centers & Institutes
- Students & Faculty
- Academic Divisions

- Alumna (BS '21)
Nayla Abney
Nayla Abney took a chance leaving New Jersey and the East Coast to come to Caltech. She helped launch the inaugural season for women's soccer at Caltech in 2017 and says the sport and the team teach lessons that help her in the classroom and on the field. The chemical engineering major is inspired by the researchers and professors on campus, and she is committed to building a legacy for other young women at Caltech.

- Alumnus (BS '20)
David Ignacio Fager
For as long as he can remember, David Ignacio Fager has adored mathematics. In high school, he lived for Mu Alpha Theta competitions and skipped ahead in math textbooks the way impatient readers sometimes peek at the last page of a mystery novel. Then came freshman year of college, and Caltech’s social science core introduced him to a new love: economics.

- Bren Professor of Psychology, Neuroscience, and Biology
Ralph Adolphs
Since his years as a Caltech graduate student, Ralph Adolphs (PhD ’93) has wanted to learn how the biological brain produces the intangible mind, what the mind’s basic elements are, and how the two influence each other.

- Assistant Professor of Planetary Science
Katherine de Kleer
Katherine de Kleer uses a diverse range of telescopes to observe planets and their moons at radio, infrared, and optical wavelengths. Her innovative approaches for studying this wide swath of frequencies have helped shed light on the seasonal evolution of planetary atmospheres.

Connect With Us
Stay up to date on the latest from Caltech.
Get the latest news from the Caltech website delivered to your email inbox.

- #Caltech2024

Caltech's Commitment to an Inclusive Environment

- New Technique Could Help Build Quantum Computers of the Future
Key Takeaways
- Berkeley Lab researchers have reported a major advancement that could bring us closer to a scalable quantum computer.
- Using a femtosecond laser during experiments which explore the role of hydrogen in qubit formation, the researchers developed a method that programs the formation of telecom-band optical qubits in silicon for large-scale manufacturing.
- The technique could enable scalable quantum computers of the future by building on current silicon-based computing infrastructure.
Quantum computers have the potential to solve complex problems in human health, drug discovery, and artificial intelligence millions of times faster than some of the world’s fastest supercomputers. A network of quantum computers could advance these discoveries even faster. But before that can happen, the computer industry will need a reliable way to string together billions of qubits – or quantum bits – with atomic precision.
Connecting qubits, however, has been challenging for the research community. Some methods form qubits by placing an entire silicon wafer in a rapid annealing oven at very high temperatures. With these methods, qubits randomly form from defects (also known as color centers or quantum emitters) in silicon’s crystal lattice. And without knowing exactly where qubits are located in a material, a quantum computer of connected qubits will be difficult to realize.
But now, getting qubits to connect may soon be possible. A research team led by Lawrence Berkeley National Laboratory (Berkeley Lab) says that they are the first to use a femtosecond laser to create and “annihilate” qubits on demand, and with precision, by doping silicon with hydrogen.
The advance could enable quantum computers that use programmable optical qubits or “spin-photon qubits” to connect quantum nodes across a remote network. It could also advance a quantum internet that is not only more secure but could also transmit more data than current optical-fiber information technologies.
“This could carve out a potential new pathway for industry to overcome challenges in qubit fabrication and quality control.” – Thomas Schenkel, senior scientist, Accelerator Technology & Applied Physics Division
“To make a scalable quantum architecture or network, we need qubits that can reliably form on-demand, at desired locations, so that we know where the qubit is located in a material. And that’s why our approach is critical,” said Kaushalya Jhuria, a postdoctoral scholar in Berkeley Lab’s Accelerator Technology & Applied Physics (ATAP) Division. She is the first author on a new study that describes the technique in the journal Nature Communications . “Because once we know where a specific qubit is sitting, we can determine how to connect this qubit with other components in the system and make a quantum network.”
“This could carve out a potential new pathway for industry to overcome challenges in qubit fabrication and quality control,” said principal investigator Thomas Schenkel , head of the Fusion Science & Ion Beam Technology Program in Berkeley Lab’s ATAP Division. His group will host the first cohort of students from the University of Hawaii in June as part of a DOE Fusion Energy Sciences-funded RENEW project on workforce development where students will be immersed in color center/qubit science and technology.
Forming qubits in silicon with programmable control
The new method uses a gas environment to form programmable defects called “color centers” in silicon. These color centers are candidates for special telecommunications qubits or “spin photon qubits.” The method also uses an ultrafast femtosecond laser to anneal silicon with pinpoint precision where those qubits should precisely form. A femtosecond laser delivers very short pulses of energy within a quadrillionth of a second to a focused target the size of a speck of dust.
Spin photon qubits emit photons that can carry information encoded in electron spin across long distances – ideal properties to support a secure quantum network. Qubits are the smallest components of a quantum information system that encodes data in three different states: 1, 0, or a superposition that is everything between 1 and 0.
With help from Boubacar Kanté, a faculty scientist in Berkeley Lab’s Materials Sciences Division and professor of electrical engineering and computer sciences (EECS) at UC Berkeley, the team used a near-infrared detector to characterize the resulting color centers by probing their optical (photoluminescence) signals.
What they uncovered surprised them: a quantum emitter called the Ci center. Owing to its simple structure, stability at room temperature, and promising spin properties, the Ci center is an interesting spin photon qubit candidate that emits photons in the telecom band. “We knew from the literature that Ci can be formed in silicon, but we didn’t expect to actually make this new spin photon qubit candidate with our approach,” Jhuria said.
An artistic depiction of a new method to create high-quality color-centers (qubits) in silicon at specific locations using ultrafast laser pulses (femtosecond, or one quadrillionth of a second). The inset at the top-right shows an experimentally observed optical signal (photoluminescence) from the qubits, with their structures displayed at the bottom. (Credit: Kaushalya Jhuria/Berkeley Lab)
The researchers learned that processing silicon with a low femtosecond laser intensity in the presence of hydrogen helped to create the Ci color centers. Further experiments showed that increasing the laser intensity can increase the mobility of hydrogen, which passivates undesirable color centers without damaging the silicon lattice, Schenkel explained.
A theoretical analysis performed by Liang Tan, staff scientist in Berkeley Lab’s Molecular Foundry, shows that the brightness of the Ci color center is boosted by several orders of magnitude in the presence of hydrogen, confirming their observations from laboratory experiments.
“The femtosecond laser pulses can kick out hydrogen atoms or bring them back, allowing the programmable formation of desired optical qubits in precise locations,” Jhuria said.
The team plans to use the technique to integrate optical qubits in quantum devices such as reflective cavities and waveguides, and to discover new spin photon qubit candidates with properties optimized for selected applications.
“Now that we can reliably make color centers, we want to get different qubits to talk to each other – which is an embodiment of quantum entanglement – and see which ones perform the best. This is just the beginning,” said Jhuria.
“The ability to form qubits at programmable locations in a material like silicon that is available at scale is an exciting step towards practical quantum networking and computing,” said Cameron Geddes, Director of the ATAP Division.
Theoretical analysis for the study was performed at the Department of Energy’s National Energy Research Scientific Computing Center (NERSC) at Berkeley Lab with support from the NERSC QIS@Perlmutter program.
The Molecular Foundry and NERSC are DOE Office of Science user facilities at Berkeley Lab.
This work was supported by the DOE Office of Fusion Energy Sciences.
Lawrence Berkeley National Laboratory (Berkeley Lab) is committed to delivering solutions for humankind through research in clean energy, a healthy planet, and discovery science. Founded in 1931 on the belief that the biggest problems are best addressed by teams, Berkeley Lab and its scientists have been recognized with 16 Nobel Prizes. Researchers from around the world rely on the Lab’s world-class scientific facilities for their own pioneering research. Berkeley Lab is a multiprogram national laboratory managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science .

- Emerging Capabilities
Building AI agents with Semantic Kernel
via InfoWorld
Jan. 25, 2024
- #artificial intelligence
- #computer science
- Pattie Maes Professor of Media Technology; Germeshausen Professor
- Media Lab Research Theme: Life with AI
Share this article
By Simon Bisson
Back in the early 1990s, I worked in a large telecoms research lab, as part of the Advanced Local Loop group. Our problem domain was the “last mile”—getting services to peoples’ homes. One of my research areas involved thinking about what might happen when the network shift from analog to digital services was complete.
I spent a great deal of time in the lab’s library, contemplating what computing would look like in a future of universal bandwidth. One of the concepts that fascinated me was ubiquitous computing, where computers disappear into the background and software agents become our proxies, interacting with network services on our behalf. That idea inspired work at Apple, IBM, General Magic, and many other companies.
One of the pioneers of the software agent concept was MIT professor Pattie Maes. Her work crossed the boundaries between networking, programming, and artificial intelligence, and focused on two related ideas: intelligent agents and autonomous agents. These were adaptive programs that could find and extract information for users and change their behavior while doing so.
It has taken the software industry more than 30 years to catch up with that pioneering research, but with a mix of transformer-based large language models (LLMs) and adaptive orchestration pipelines, we’re finally able to start delivering on those ambitious original ideas.
ACM ByteCast: Pattie Maes
On the ACM ByteCast podcast, Fluid Interfaces head Pattie Maes talks to host Rashmi Mohan about the passion that has shaped her career.

Research Group Overview: Fluid Interfaces
Designing systems for cognitive support
Pattie Maes reaches 500 peer-reviewed accepted papers
Pattie Maes, head of the Fluid Interfaces group and Professor of Media Technology, reaches milestone of 500 peer-reviewed accepted papers.
Interaction with Embodied Media
Merrill, D. "Interaction with Embodied Media"

- B.S. Students
- M.S. Students
- Ph.D. Students
- D-Clearance
- Directed Research
- Information for Graders and Course Producers
- Microsoft Imagine
- CS Student Organizations
- CS Library Guide
- CS Job Announcements
[UG/MS/PhD] ACE Lab Research Study Recruitment
![research about computer laboratory Featured image for “[UG/MS/PhD] ACE Lab Research Study Recruitment”](https://www.cs.usc.edu/wp-content/uploads/2024/04/USC.png)
The following announcement is from Xinyi Zhou ([email protected]). Please contact them directly if you have any questions.
Hi everyone!
We at the Adaptive Computing Experiences (ACE) Lab at the University of Southern California are conducting a research study to investigate biases in Human-AI Software Development Teams. We would love to hear your thoughts and experiences!
I am recruiting individuals who meet the following criteria for the study:
- 18 years old or above
- Reside in the United States
- English Speaker
- Programming experience of at least three years for students
- Regularly uses Large Language Model (LLM) tools for software development
If you decide to participate in this study, you will be asked to do the following activities:
- Complete a quick online survey.
- Participate in a 1:1 online session for 60 minutes.
- Participate in a 15-20-minute 1:1 interview after the online session.
During these activities, you will be asked questions about:
- Demographic details include age, occupation, role, years of experience, etc.
- Exploration of Cognitive Biases Related Behaviors and Their Impacts
Participants in this study will receive a $40 reward after the final interview.
If you are interested in participating in this study, please click this link to fill out our survey. If you have questions, please contact me at [email protected].
Published on June 4th, 2024
Last updated on June 4th, 2024
- CS Announcements
- Job/Research Opportunities
- Undergraduate
- Chair’s Welcome
- Awards and Honors
- CS@SC Institutes
- Media Coverage
- Newsletters and Fact Sheets
- CS Industry Affiliate Program
- Bekey Lecture
- Driving Directions
- Open Staff Positions
- Open Faculty Positions
- Centers and Institutes
- Research Areas and Labs
- Technical Reports
- Annual Research Review
- Undergraduate Research Experiences
- Faculty Directory
- Staff Directory
- Getting Started with CS@USC
- B.S. Program
- M.S. Program
- Ph.D. Program
- Data Science Program
- Graduate Certificate
- Distance Education
- K-12 Outreach
- Academic Advisement
- B.S. Application Information
- M.S. Application Information
- Ph.D. Application Information

IMAGES
VIDEO
COMMENTS
18. QUALITY COMP UTER LABS PROMOTE STUDENT SUCCESS. Adnan Omar and Muhammed Miah*. Department of Computer Information Systems, Southern Universi ty at New Orleans, 6801. Press Drive, New Orleans ...
The increasing use of personal and portable technologies such as smartphones and tablets, together with the availability of fabrication technologies, has led to recent reenvisioning of the computer laboratory as a space where digital technologies are used as part of integrated research, design, and realization activities.
The Computer Laboratory Environment Inventory (CLEI) and The Attitude Towards Computer and Computer Courses (ACCC) s Fisher, 1998). The sample consisted of 250 students taken from private ...
Based on the research that has been done, it’s concluded that: (1) description of computer laboratory facilities, students’ learning interest and students’ mathematics ...
The Computer Science and Artificial Intelligence Laboratory (CSAIL) pursues fundamental research across the entire breadth of computer science and artificial intelligence. CSAIL is committed to leading the field both in new theoretical approaches and in the creation of applications that have broad societal impact.
Research. The computing and information revolution is transforming society. Cornell Computer Science is a leader in this transformation, producing cutting-edge research in many important areas. The excellence of Cornell faculty and students, and their drive to discover and collaborate, ensure our leadership will continue to grow.
This study focuses on the computer laboratory class as a learning environment in university courses. It involved the development and validation of two instruments, the Computer Laboratory Environment Inventory (CLEI) and the Attitude towards Computing and Computing Courses Questionnaire (ACCC). The CLEI has five scales for measuring students ...
Computer-Assisted Medicine. Computer science research at the Johns Hopkins University is advancing computing technology, enabling new modes of thought, and transforming society. Our faculty conduct innovative, collaborative research aimed at solving large and complex interdisciplinary problems, drawing upon the university's renowned strengths ...
A computer laboratory is an expensive resource in terms of equipment and people, and should be used as effectively as possible. Computer laboratory classes may be organized as closed laboratories which are scheduled and staffed in the same way as other classes, or as open laboratories where students come and go as they please.
Six tools group leaders love. • AnyDesk is free software for accessing and controlling computers remotely. • Google's Apps Script automates actions across the Google application suite ...
Developed computer laboratory will benefit the students in research. 2. Undeveloped computer laboratory will not benefit the students in research "Improving the quality of computer labortatory for Senior High School students of Mount Carmel School of Maria Aurora Inc. (MCSMA): An Evaluation" 6 MOUNT CARMEL SCHOOL OF MARIA AURORA, (MCSMA) INC.
About. USC has a strong and active background in modern theoretical computer science, with research spanning a broad range of topics. Areas of particular interest include the theory of algorithms and optimization, graph theory, scalable algorithms, theory of machine learning, computational geometry, complex analysis, computational complexity, algorithmic number theory and cryptography.
Computer Science and Artificial Intelligence Laboratory (CSAIL) The largest interdepartmental laboratory at MIT that focuses on developing fundamental new technologies, conducting basic research that furthers the field of computing, and inspiring and educating future generations of scientists and technologists. View CSAIL.
The Computer Architecture (comparch) Lab conducts research on all aspects of future microprocessor technology including performance, power, multi-threading, chip-multiprocessing, security, programmability, reliability, interaction with compilers and software, and the impact of future technologies. Data Systems and Analytics Group
The Computer Architecture lab is dedicated to the engineering of machines that learn faster, compute more efficiently, operate safer, program easier, and integrate more responsibly. It aims to be an inviting and interdisciplinary space to rethink what a computer system can be. ... The group's research interests are in computer systems and ...
Welcome to the Computer Systems Lab (CSL) at Yale University.. CSL is an interdisciplinary laboratory with faculty from both Electrical Engineering and Computer Science that have a shared research interest in computer systems. CSL research encompasses both theory and practice in a wide range of domains including architecture, biologically-inspired computing, certified systems software ...
Massachusetts Institute of Technology. Computer Science & Artificial Intelligence Laboratory. 32 Vassar St, Cambridge MA 02139
The ELX lab conducts research in human-computer interaction, with a focus on learning technologies and technologies to support mental wellness. Research areas include wearable technologies for learning, enactment-based storytelling, Maker technologies in education and narrative-centered technological approaches. The goal of the lab is to design ...
Machina and Wild investigated the importance of ELNs when integrated with other computer tools, such as laboratory information management systems, analytical instrumentation, data management systems, and scientific data. They observed that the type of laboratory (analytical, synthesis, clinical, research) was a primary source of differences ...
Mission: The VLM lab is a research lab at the Computer Science and Engineering Department of the University of Texas at Arlington. At the VLM lab we are conducting research in the areas of computer vision, machine learning, and data mining. Areas of focus include gesture and sign language recognition, human motion analysis, detection and ...
The Prescience Lab conducts a range of experimental computer systems research with a current focus on virtualization, empathic systems, and parallel and distributed systems. This lab is headed by Peter Dinda. ... The VLSI Research Lab at NU focuses on the areas of energy efficient computing with digital and mixed signal design approaches. We ...
Research Labs. Automated Reasoning Group (Adnan Darwiche) Big Data and Genomics Lab (Eran Halperin) Biocybernetics Laboratory (Joe DiStefano) Center for Smart Health (Ramin Ramezani) Center for Vision, Cognition, Learning and Autonomy (Song-Chun Zhu) Cognitive Systems Laboratory (Judea Pearl) Computational Genetics Laboratory (Eleazar Eskin ...
The Computer Systems Research Laboratory is a research group made up of faculty members and students in the Department of Computer Science and Engineering at the University of North Texas. While we are interested in many topics within computer science, our current research focuses mainly on 3D RAM design, Processing-in-Memory and memory ...
The University of the Philippines (UP) Computer Security Group (CSG) is a research laboratory established in 2002. It is focused on the study of security mechanisms and protocols with a particular emphasis in the design and development of secure applications. As part of its affi….
Li's co-authors include Ruonan Han, an associate professor in EECS, leader of the Terahertz Integrated Electronics Group, and member of the Research Laboratory of Electronics (RLE); senior author Dirk Englund, professor of EECS, principal investigator of the Quantum Photonics and Artificial Intelligence Group and of RLE; as well as others at ...
Enhancing engineering education through mini project-based learning in computer integrated manufacturing laboratory: A student-centric approach. Ravikantha Prabhu ... holds a Master's degree from Srinivas Institute of Technology and a Ph.D. from VTU, Belagavi. His research focused on biodiesel with aqueous nanoparticles. His interests include ...
Caltech does not discriminate or permit discrimination by any member of its community on the basis of sex, race, color, religion, national origin, citizenship, ancestry, age, marital status, physical or mental disability, medical condition, genetic information, pregnancy or perceived pregnancy, gender, gender identity or expression, sexual orientation, protected veteran status, or any other ...
A research team led by Lawrence Berkeley National Laboratory (Berkeley Lab) says that they are the first to use a femtosecond laser to create and "annihilate" qubits on demand, and with precision, by doping silicon with hydrogen. The advance could enable quantum computers that use programmable optical qubits or "spin-photon qubits" to ...
Building AI agents with Semantic Kernel. By Simon Bisson. Back in the early 1990s, I worked in a large telecoms research lab, as part of the Advanced Local Loop group. Our problem domain was the "last mile"—getting services to peoples' homes. One of my research areas involved thinking about what might happen when the network shift from ...
The following announcement is from Xinyi Zhou ([email protected]). Please contact them directly if you have any questions. Hi everyone! We at the Adaptive Computing Experiences (ACE) Lab at the University of Southern California are conducting a research study to investigate biases in Human-AI Software Development Teams. We would love to hear your thoughts and experiences! I am recruiting ...